Getting started
Gao Wang, Anjing Liu and William Denault
Source:vignettes/getting_started.Rmd
getting_started.RmdmfsusieR is the multi-outcome, functional regression
generalization of susieR::susie(). It provides two main
functions:
-
fsusie(Y, X, pos = NULL, ...)for fine-mapping a single response.Ymay be scalar (T = 1, recoverssusieR::susie()) or functional (T > 1, the single-outcome functional SuSiE model). -
mfsusie(X, Y, pos = NULL, ...)for fine-mapping multiple responses jointly.Yis a list of lengthMoutcomes; each element is a matrixn x T_m(withT_m = 1for scalar outcomes).
Both functions return an object of class
c("mfsusie", "susie") with the standard SuSiE fields
(alpha, mu, mu2,
lbf, KL, sigma2,
elbo, pip, sets,
niter, converged).
Both fits return raw inverse-DWT effect curves through
coef(). For a smoothed position-space curve with a
pointwise credible band (and, for the HMM smoother, a per-position lfsr
curve), post-process with mf_post_smooth(). Four methods
are available — "TI" (the default; translation-invariant
wavelet denoising via cycle spinning), "scalewise"
(per-scale soft- thresholding), "HMM" (hidden Markov model
with per-position lfsr), and "smash" (delegates to
smashr::smash.gaus). See vignette("post_processing")
for the walk-through and the comparison workflow when several smoothers
coexist on one fit.
A small fsusie() example (single outcome,
functional)
Simulate one functional response on a length-32 grid
with two causal variables.
n <- 200; p <- 50; T_m <- 32
X <- matrix(rnorm(n * p), nrow = n)
beta <- numeric(p); beta[c(5, 17)] <- c(1.2, -0.8)
Y <- X %*% matrix(rep(beta, T_m), nrow = p) +
matrix(rnorm(n * T_m, sd = 0.4), nrow = n)
fit_f <- fsusie(Y, X, L = 15, L_greedy = 5, verbose = TRUE)
#> iter ELBO delta sigma2 mem V extras
#> 1 -783.1957 - [0.059, 0.064, 30.044] 0.16 GB [1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] pi_null=[1.00, 1.00]
#> iter 2: max|dPIP|=9.80e-01, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.16 GB]
#> iter 3: max|dPIP|=0.00e+00, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] -- converged [mem: 0.16 GB]
#> [L_greedy] 1 round, greedy_lbf_cutoff=0.100, final L=5
#> round L min(lbf) action
#> 1 5 0.000 saturated
fit_f$pip[c(5, 17)]
#> [1] 1 1
length(fit_f$sets$cs)
#> [1] 2fsusie() is a wrapper around
mfsusie(X, list(Y), list(pos), ...) with no numerical
difference. mfsusie_plot() works on the result either
way:
plot_dims_f <- mfsusie_plot_dimensions(fit_f)
mfsusie_plot(fit_f)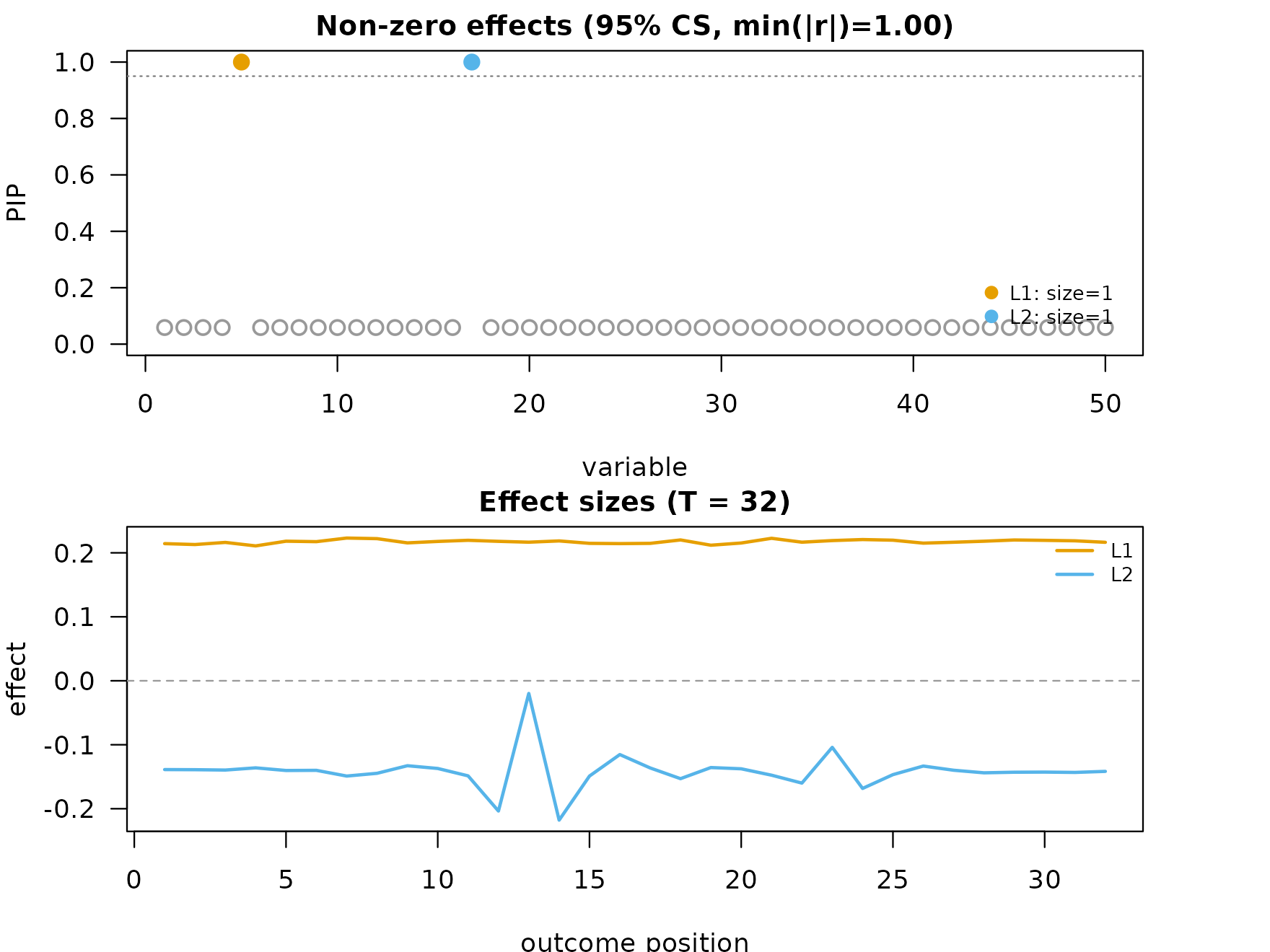
A small mfsusie() example (multi-outcome)
Same X and effect vector, two functional outcomes at
different resolutions plus one scalar outcome.
T_per <- c(32L, 64L)
Y_func <- lapply(T_per, function(T_m) {
X %*% matrix(rep(beta, T_m), nrow = p) +
matrix(rnorm(n * T_m, sd = 0.4), nrow = n)
})
Y_scalar <- as.numeric(X %*% beta + rnorm(n, sd = 0.4))
Y_list <- c(Y_func, list(matrix(Y_scalar, ncol = 1)))
fit_m <- mfsusie(X, Y_list, L = 15, L_greedy = 5,
prior_variance_scope = "per_outcome",
verbose = TRUE)
#> iter ELBO delta sigma2 mem V extras
#> 1 -2387.0080 - [0.050, 0.064, 60.004] 0.16 GB [1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] pi_null=[1.00, 1.00]
#> iter 2: max|dPIP|=1.45e-08, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] -- converged [mem: 0.16 GB]
#> [L_greedy] 1 round, greedy_lbf_cutoff=0.100, final L=5
#> round L min(lbf) action
#> 1 5 0.000 saturated
fit_m$pip[c(5, 17)]
#> [1] 1 1
length(fit_m$sets$cs)
#> [1] 2mu[[l]][[m]] is a p x T_basis[m] matrix of
per-effect posterior means on each outcome; predict() and
coef() invert the wavelet transform to position space
transparently. mfsusie_plot() tiles one effect-curve panel
per outcome with the PIPs in the top-left slot:
plot_dims_m <- mfsusie_plot_dimensions(fit_m)
mfsusie_plot(fit_m)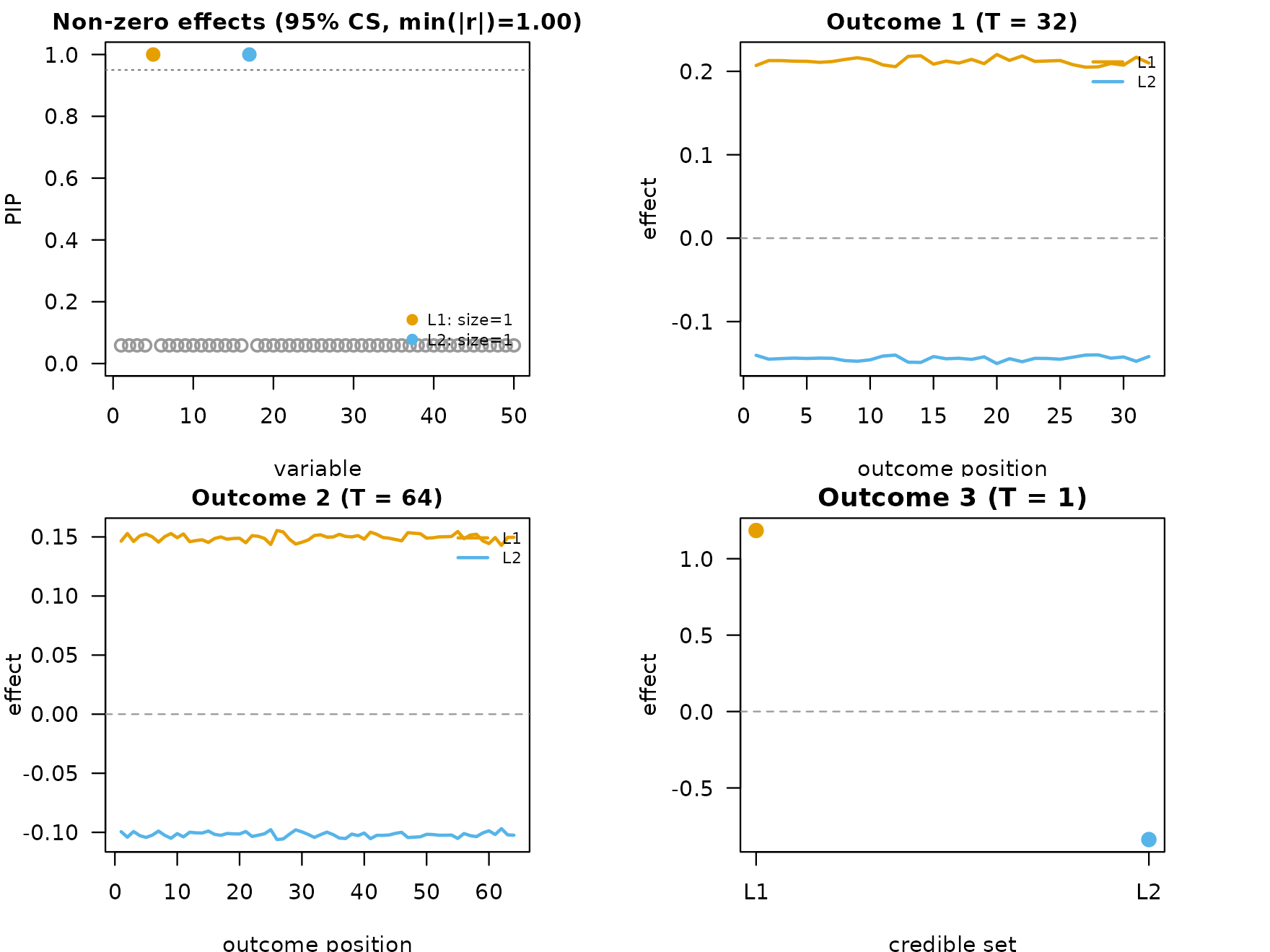
coef(fit_m) returns a per-outcome list of
L x T_basis[m] matrices on the original X scale: row
l of element m is the inverse-DWT’d effect
curve for effect l on outcome m.
cf <- coef(fit_m)
length(cf) # one per outcome (M)
#> [1] 3
dim(cf[[1L]]) # L x T_basis[1]
#> [1] 5 32predict(fit_m, newx = X_new) returns the fitted
per-position response on a fresh genotype matrix:
X_new <- matrix(rnorm(20 * p), 20, p)
pr <- predict(fit_m, newx = X_new)
length(pr); dim(pr[[1L]]) # n_new x T_basis[m]
#> [1] 3
#> [1] 20 32
# Plot the predicted curve for the first outcome on the first
# four held-out samples.
matplot(t(pr[[1L]][seq_len(4L), ]), type = "l", lty = 1,
xlab = "position", ylab = "predicted Y",
main = "predict.mfsusie() on outcome 1")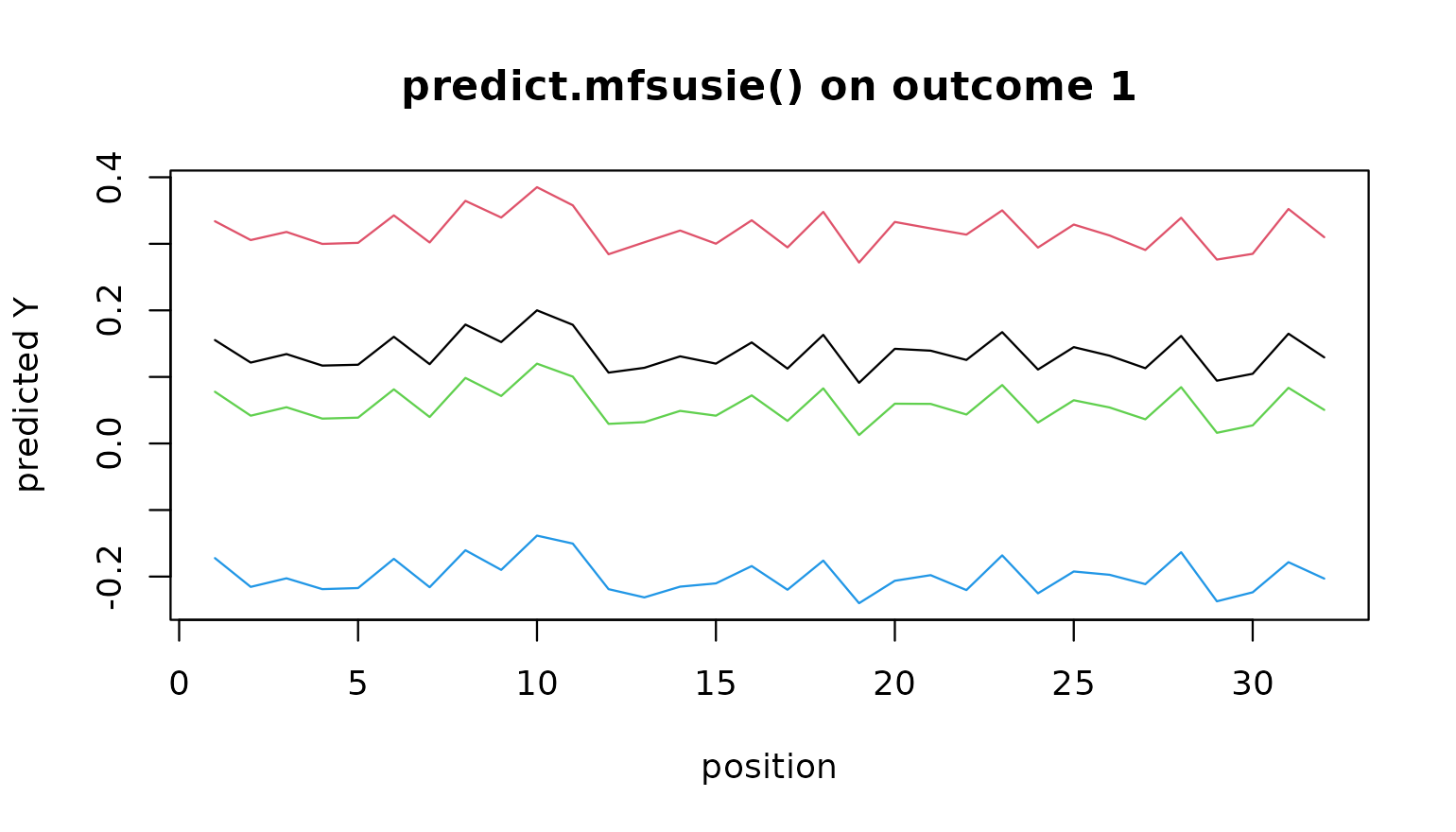
Inspecting a fit
summary() prints a one-screen overview of any
mfsusie / fsusie fit: dimensions, IBSS
iterations and convergence, the final ELBO, the credible-set table
(per-CS size, purity, lead SNP), top PIPs, and the mixture-prior
null-mass summary across all (outcome, scale) cells.
summary(fit_f)
#> mfsusie summary: p=50, L=5, M=1, converged in 3 iter
#> T_basis per outcome: (32)
#> Final ELBO: -240.8975
#> Mixture null-mass across (m, s): min=1.000, median=1.000, max=1.000
#> Credible sets:
#> CS 1: size=1, purity=1.000, variables=5
#> CS 2: size=1, purity=1.000, variables=17The same call works on the multi-outcome fit and shows the same
structure with the per-outcome T_basis:
summary(fit_m)
#> mfsusie summary: p=50, L=5, M=3, converged in 2 iter
#> T_basis per outcome: (32, 64, 1)
#> Final ELBO: -18296.6506
#> Mixture null-mass across (m, s): min=1.000, median=1.000, max=1.000
#> Credible sets:
#> CS 1: size=1, purity=1.000, variables=5
#> CS 2: size=1, purity=1.000, variables=17Numerical equivalence with susieR::susie
When Y is scalar, the prior collapses to a single
Gaussian component, the null component is removed, and the per-(scale,
outcome) variance structure collapses to a scalar,
mfsusie() reduces exactly to
susieR::susie() when we set it a fixed effects size
prior.
y_scalar <- as.numeric(X %*% beta + rnorm(n, sd = 0.4))
y <- (y_scalar - mean(y_scalar)) / sd(y_scalar)
fit_s <- susie(X, y, L = 5,
scaled_prior_variance = 0.2,
estimate_prior_variance = FALSE,
estimate_residual_variance = TRUE,
max_iter = 100, tol = 1e-8)
fit_d <- mfsusie(X, list(matrix(y, ncol = 1)),
L = 5,
prior_variance_grid = 0.2,
prior_variance_scope = "per_outcome",
null_prior_weight = 0,
residual_variance_scope = "per_outcome",
estimate_prior_variance = FALSE,
L_greedy = NULL,
max_iter = 100, tol = 1e-8,
verbose = FALSE)
# Element-wise agreement on every numeric field.
max(abs(fit_d$alpha - fit_s$alpha))
#> [1] 1.930582e-06
max(abs(fit_d$pip - fit_s$pip))
#> [1] 1.279103e-06
max(abs(fit_d$sigma2[[1]] - fit_s$sigma2))
#> [1] 8.596112e-09
max(abs(fit_d$lbf - fit_s$lbf))
#> [1] 0.000248286
max(abs(fit_d$KL - fit_s$KL))
#> [1] 3.134916e-06
abs(tail(fit_d$elbo, 1) - tail(fit_s$elbo, 1))
#> [1] 3.900169e-11
identical(fit_d$niter, fit_s$niter)
#> [1] FALSEThe credible-set memberships are also identical:
This degenerate-case is introduced as a sanity check, as
mfsusieR is implemented on the backbone of
susieR and is expect to produce the same result
mathematically under degenerated-case like this.
Where to next
- Single-outcome applications: Single-outcome intro, covariate adjustment + colocalization, DNAm case study, why functional.
- Multi-outcome applications: Multi-outcome intro, long-running fits.
Session info
This is the version of R and the packages that were used to generate these results.
sessionInfo()
#> R version 4.4.3 (2025-02-28)
#> Platform: x86_64-conda-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS/LAPACK: /home/runner/work/mfsusieR/mfsusieR/.pixi/envs/r44/lib/libopenblasp-r0.3.32.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: Etc/UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] susieR_0.16.1 mfsusieR_0.0.1
#>
#> loaded via a namespace (and not attached):
#> [1] sass_0.4.10 generics_0.1.4 ashr_2.2-63
#> [4] lattice_0.22-9 magrittr_2.0.5 digest_0.6.39
#> [7] evaluate_1.0.5 grid_4.4.3 RColorBrewer_1.1-3
#> [10] fastmap_1.2.0 plyr_1.8.9 jsonlite_2.0.0
#> [13] Matrix_1.7-5 reshape_0.8.10 mixsqp_0.3-54
#> [16] scales_1.4.0 truncnorm_1.0-9 invgamma_1.2
#> [19] textshaping_1.0.5 jquerylib_0.1.4 cli_3.6.6
#> [22] rlang_1.2.0 zigg_0.0.2 crayon_1.5.3
#> [25] LaplacesDemon_16.1.8 cachem_1.1.0 yaml_2.3.12
#> [28] otel_0.2.0 tools_4.4.3 SQUAREM_2026.1
#> [31] parallel_4.4.3 dplyr_1.2.1 wavethresh_4.7.3
#> [34] ggplot2_4.0.3 Rfast_2.1.5.2 vctrs_0.7.3
#> [37] R6_2.6.1 matrixStats_1.5.0 lifecycle_1.0.5
#> [40] fs_2.1.0 htmlwidgets_1.6.4 MASS_7.3-65
#> [43] ragg_1.5.2 irlba_2.3.7 pkgconfig_2.0.3
#> [46] desc_1.4.3 pillar_1.11.1 pkgdown_2.2.0
#> [49] RcppParallel_5.1.11-2 bslib_0.10.0 gtable_0.3.6
#> [52] glue_1.8.1 Rcpp_1.1.1-1.1 systemfonts_1.3.2
#> [55] tidyselect_1.2.1 tibble_3.3.1 xfun_0.57
#> [58] knitr_1.51 dichromat_2.0-0.1 farver_2.1.2
#> [61] htmltools_0.5.9 rmarkdown_2.31 compiler_4.4.3
#> [64] S7_0.2.2