Two-pass scans with warm-start follow-up
Gao Wang, Anjing Liu and William Denault
Source:vignettes/mfsusie_long_running_fits.Rmd
mfsusie_long_running_fits.RmdJoint multi-outcome fits over many ATAC-seq conditions, dense
position grids, or large p (number of predictors) can run
beyond the wall time of a typical cluster job. Genome-wide scans
compound the problem: most loci carry no fine-mapping signal, yet a
uniform iteration budget spends the same effort on them as on the loci
that actually need it.
The pattern this vignette covers is a two-pass scan:
-
First pass: run every locus with a low
max_iterbudget (e.g. 5). Two outcomes are useful directly. Loci that converge withconverged = TRUEare done. Loci that did not converge but whose first-pass PIPs are flat (no SNP near 1, no narrow credible set) are unlikely to produce signal at any budget and can also be dropped. Filtering on these criteria alone usually drops the bulk of a genome-wide scan. - Second pass: re-fit the remaining candidates (didn’t converge, but first-pass PIPs already concentrate on a small number of SNPs) with a larger budget, warm-started from the partial fit so the IBSS loop does not restart from cold and just polishes the partial state.
Setup: one signal locus and one null locus
data(N3finemapping)
X <- N3finemapping$X[, seq_len(120)]
n <- nrow(X); p <- ncol(X)
T_func <- c(32L, 32L)
# Locus A: real cis-signal at SNPs 37 and 88.
beta_signal <- numeric(p); beta_signal[c(37L, 88L)] <- c(1, -0.6)
Y_signal <- lapply(T_func, function(T_m) {
X %*% matrix(rep(beta_signal, T_m), nrow = p) +
matrix(rnorm(n * T_m, sd = 0.4), nrow = n)
})
# Locus B: no signal (independent noise).
Y_null <- lapply(T_func, function(T_m)
matrix(rnorm(n * T_m, sd = 0.4), nrow = n))First pass: short budget, screen everything
max_iter = 5 is enough for null loci to settle and for
signal loci to expose themselves through converged = FALSE
plus an ELBO that is still rising. We use track_fit = TRUE
only to inspect the trajectory; in production it is optional.
fit_signal_short <- mfsusie(X, Y_signal, L = 15, L_greedy = 5,
max_iter = 5,
prior_variance_scope = "per_outcome",
verbose = FALSE)
fit_null_short <- mfsusie(X, Y_null, L = 15, L_greedy = 5,
max_iter = 5,
prior_variance_scope = "per_outcome",
verbose = FALSE)
c(signal = fit_signal_short$niter, null = fit_null_short$niter)
#> signal null
#> 4 2
c(signal = fit_signal_short$converged, null = fit_null_short$converged)
#> signal null
#> TRUE TRUE
length(fit_signal_short$sets$cs)
#> [1] 2
length(fit_null_short$sets$cs)
#> [1] 0The null locus converges in a couple of iterations and produces no
credible sets. The signal locus reports converged = FALSE
at the budget — its ELBO is still rising — and is the candidate to
follow up on.
Triage: drop converged-no-signal, queue non-converged-with-signal
loci <- list(signal = fit_signal_short, null = fit_null_short)
followup <- vapply(loci, function(f) {
!isTRUE(f$converged) || length(f$sets$cs) > 0L
}, logical(1L))
followup
#> signal null
#> TRUE FALSEOnly the signal locus is queued for the second pass. A production scan would loop this triage over all loci and emit the followup queue.
Second pass: warm-start from the partial fit
model_init accepts a previously returned mfsusie fit.
The IBSS loop seeds alpha, mu,
mu2, V, pi_V,
G_prior, sigma2, and fitted from
the supplied object instead of the cold zero state. The follow-up then
converges in a small number of additional iterations rather than the
cold-start total.
fit_signal_warm <- mfsusie(X, Y_signal, L = 15, L_greedy = 5,
max_iter = 100,
model_init = fit_signal_short,
prior_variance_scope = "per_outcome",
verbose = FALSE)
fit_signal_warm$niter
#> [1] 2
fit_signal_warm$converged
#> [1] TRUECompare to a cold-start run of the same locus at full budget:
fit_signal_cold <- mfsusie(X, Y_signal, L = 15, L_greedy = 5,
max_iter = 100,
prior_variance_scope = "per_outcome",
verbose = FALSE)
fit_signal_cold$niter
#> [1] 4The cold run converges at iteration
fit_signal_cold$niter; the warm follow-up adds only the
increment between the partial budget and convergence, so wall time
scales with the residual gap rather than with the full budget.
Save and restore
Save the fit object between jobs. Re-running mfsusie with the
identical (X, Y, L, prior, seed) reproduces the same
trajectory deterministically.
saveRDS(fit_signal_short, "fit_partial.rds")
# ...later, on a new job...
fit_partial <- readRDS("fit_partial.rds")
fit_full <- mfsusie(X, Y_signal, L = 15, L_greedy = 5,
max_iter = 100,
model_init = fit_partial,
prior_variance_scope = "per_outcome")The supplied model_init must have the same
L as the new call, or fewer (in which case the warm-loaded
effects are expanded with zero-state slots up to the requested
L, the same growth pattern used by L_greedy
between rounds).
Verify the warm-start matches the cold-start fit
Two checks: the fitted posteriors should agree, and the warm trajectory should land on the same ELBO as the cold trajectory.
# Posterior agreement: PIPs, credible sets, final ELBO.
max_pip_diff <- max(abs(fit_signal_warm$pip - fit_signal_cold$pip))
elbo_diff <- abs(tail(fit_signal_warm$elbo, 1L) -
tail(fit_signal_cold$elbo, 1L))
cs_warm <- vapply(fit_signal_warm$sets$cs,
function(idx) idx[which.max(fit_signal_warm$pip[idx])],
integer(1L))
cs_cold <- vapply(fit_signal_cold$sets$cs,
function(idx) idx[which.max(fit_signal_cold$pip[idx])],
integer(1L))
data.frame(
max_pip_diff = max_pip_diff,
elbo_diff = elbo_diff,
cs_warm_leads = paste(sort(cs_warm), collapse = ","),
cs_cold_leads = paste(sort(cs_cold), collapse = ",")
)
#> max_pip_diff elbo_diff cs_warm_leads cs_cold_leads
#> 1 2.451653e-08 6.184564e-10 37,88 37,88The PIPs match within numerical noise, the final ELBOs agree, and the credible-set lead variants are identical. The warm-started run reaches the same fixed point.
# Overlay the ELBO trajectories. The warm trajectory continues
# from where the partial fit left off (iteration `n_partial + 1`).
n_partial <- fit_signal_short$niter
elbo_cold <- fit_signal_cold$elbo
elbo_warm <- c(fit_signal_short$elbo, fit_signal_warm$elbo[-1L])
xrng <- c(1, max(length(elbo_cold), length(elbo_warm)))
yrng <- range(c(elbo_cold, elbo_warm), na.rm = TRUE)
plot(seq_along(elbo_cold), elbo_cold, type = "b", pch = 19L,
col = "grey30", lwd = 2, xlim = xrng, ylim = yrng,
xlab = "iteration", ylab = "ELBO",
main = "Cold-start vs warm-start trajectories",
cex.main = 1.05, font.main = 2L)
lines(seq_along(elbo_warm), elbo_warm, type = "b", pch = 17L,
col = "firebrick", lwd = 2)
abline(v = n_partial + 0.5, lty = 3, col = "grey50")
legend("bottomright",
legend = c("cold start (full run)",
paste0("warm continuation (partial+",
fit_signal_warm$niter, " iters)"),
"partial-fit boundary"),
col = c("grey30", "firebrick", "grey50"),
lwd = c(2, 2, 1), lty = c(1, 1, 3),
pch = c(19, 17, NA), bty = "n", cex = 0.8)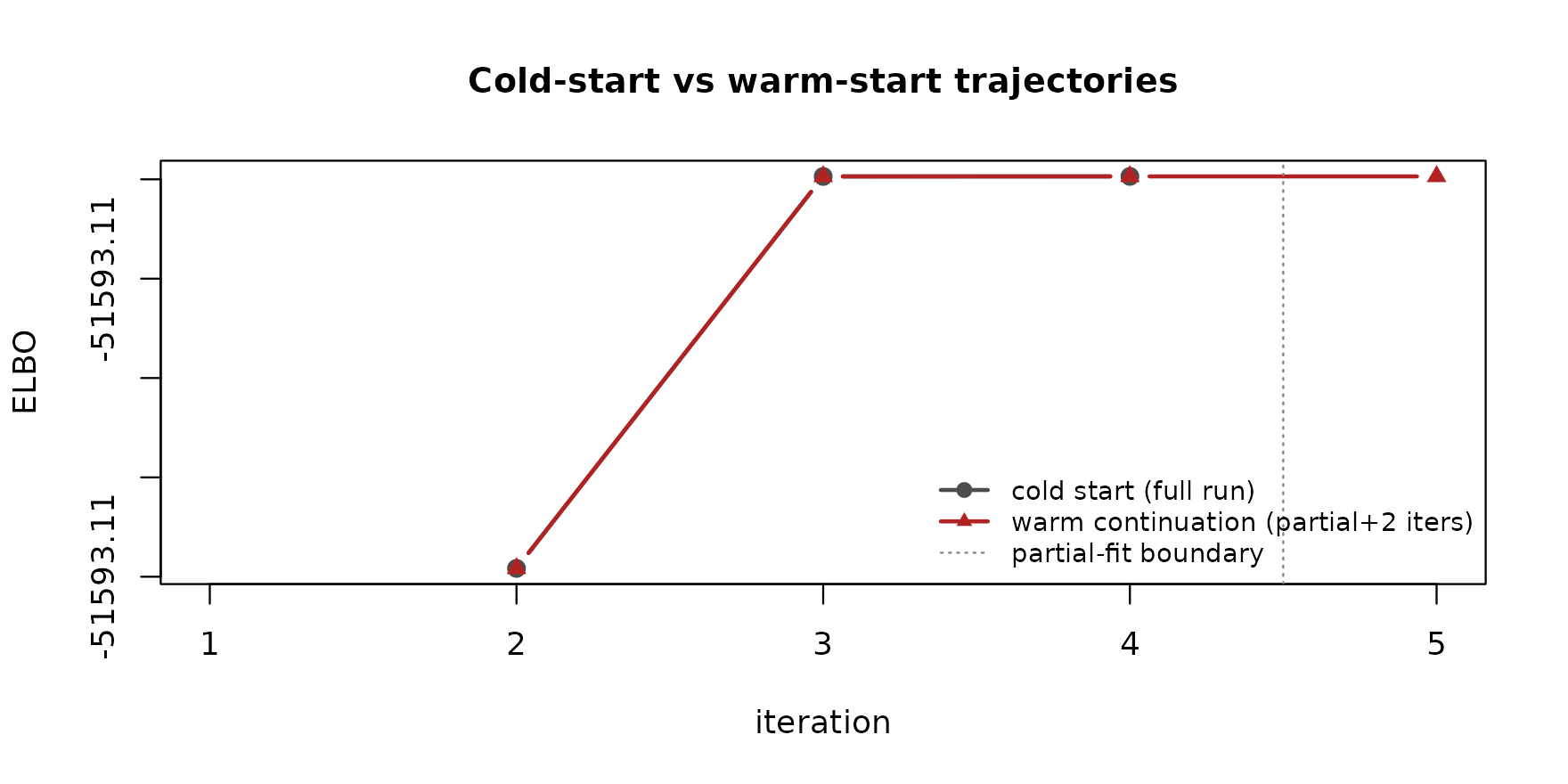
Both trajectories converge to the same ELBO. The dashed line marks where the first-pass partial fit ended; the warm continuation needs only a small increment past that point.
Diagnostic tracing inside a long run
For loci that resist convergence, track_fit = TRUE keeps
a per-iteration snapshot list at fit$trace. Each entry
records alpha (the per-effect SNP posterior,
L x p), sigma2 (the per-outcome residual
variance), pi_V (the mixture weights), and the iteration’s
ELBO.
fit_traced <- mfsusie(X, Y_signal, L = 15, L_greedy = 5,
max_iter = 20, tol = 1e-3,
prior_variance_scope = "per_outcome",
verbose = FALSE, track_fit = TRUE)
length(fit_traced$trace)
#> [1] 3
str(fit_traced$trace[[1L]], max.level = 1L)
#> List of 4
#> $ alpha : num [1:5, 1:120] 0.00833 0.00833 0.00833 0.00833 0.00833 ...
#> $ sigma2:List of 2
#> $ pi_V :List of 2
#> $ elbo : num -Inf
# How quickly does the per-iteration posterior at each true
# causal SNP settle?
true_snps <- c(37L, 88L)
alpha_path <- vapply(seq_along(fit_traced$trace), function(it) {
alpha_it <- fit_traced$trace[[it]]$alpha # L x p
vapply(true_snps, function(j) max(alpha_it[, j]),
numeric(1L))
}, numeric(2L))
rownames(alpha_path) <- paste0("SNP ", true_snps)
par(mfrow = c(1L, 2L), mar = c(4, 4, 2.5, 1))
plot(seq_len(ncol(alpha_path)), alpha_path["SNP 37", ],
type = "b", pch = 19L, col = "#1f78b4", lwd = 2,
ylim = c(0, 1),
xlab = "iteration",
ylab = "max alpha at SNP 37 across effects",
main = "Posterior trajectory at SNP 37",
cex.main = 1.05, font.main = 2L)
abline(h = 1, lty = 3, col = "grey50")
plot(seq_len(ncol(alpha_path)), alpha_path["SNP 88", ],
type = "b", pch = 19L, col = "#33a02c", lwd = 2,
ylim = c(0, 1),
xlab = "iteration",
ylab = "max alpha at SNP 88 across effects",
main = "Posterior trajectory at SNP 88",
cex.main = 1.05, font.main = 2L)
abline(h = 1, lty = 3, col = "grey50")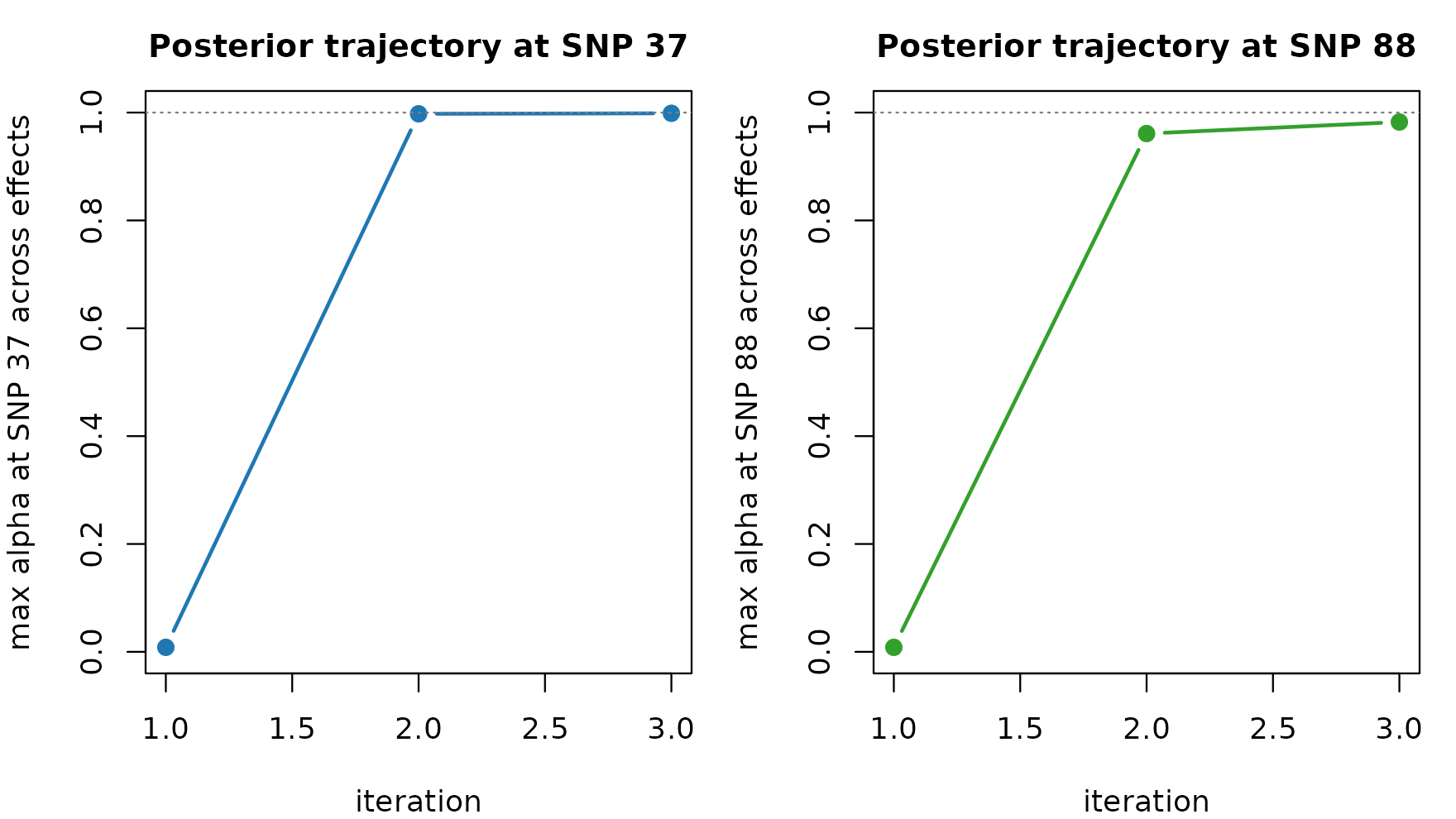
fit$trace makes it possible to see which effect
lands on which SNP at each iteration. When the IBSS loop is
slow to converge, looking at the alpha trajectories at candidate SNPs
exposes whether the loop is oscillating between competing configurations
or simply tightening posterior mass on a single one.
track_fit = TRUE is memory-heavy on real data (each
snapshot copies alpha, sigma2,
pi_V); reserve it for diagnostic runs rather than the
genome-wide screen pass. ## Session info
This is the version of R and the packages that were used to generate these results.
sessionInfo()
#> R version 4.4.3 (2025-02-28)
#> Platform: x86_64-conda-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS/LAPACK: /home/runner/work/mfsusieR/mfsusieR/.pixi/envs/r44/lib/libopenblasp-r0.3.32.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: Etc/UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] susieR_0.16.1 mfsusieR_0.0.2
#>
#> loaded via a namespace (and not attached):
#> [1] sass_0.4.10 generics_0.1.4 ashr_2.2-63
#> [4] lattice_0.22-9 magrittr_2.0.5 digest_0.6.39
#> [7] evaluate_1.0.5 grid_4.4.3 RColorBrewer_1.1-3
#> [10] fastmap_1.2.0 plyr_1.8.9 jsonlite_2.0.0
#> [13] Matrix_1.7-5 reshape_0.8.10 mixsqp_0.3-54
#> [16] scales_1.4.0 truncnorm_1.0-9 invgamma_1.2
#> [19] textshaping_1.0.5 jquerylib_0.1.4 cli_3.6.6
#> [22] rlang_1.2.0 zigg_0.0.2 crayon_1.5.3
#> [25] LaplacesDemon_16.1.8 cachem_1.1.0 yaml_2.3.12
#> [28] otel_0.2.0 tools_4.4.3 SQUAREM_2026.1
#> [31] parallel_4.4.3 dplyr_1.2.1 wavethresh_4.7.3
#> [34] ggplot2_4.0.3 Rfast_2.1.5.2 vctrs_0.7.3
#> [37] R6_2.6.1 matrixStats_1.5.0 lifecycle_1.0.5
#> [40] fs_2.1.0 htmlwidgets_1.6.4 MASS_7.3-65
#> [43] ragg_1.5.2 irlba_2.3.7 pkgconfig_2.0.3
#> [46] desc_1.4.3 pillar_1.11.1 pkgdown_2.2.0
#> [49] RcppParallel_5.1.11-2 bslib_0.10.0 gtable_0.3.6
#> [52] glue_1.8.1 Rcpp_1.1.1-1.1 systemfonts_1.3.2
#> [55] tidyselect_1.2.1 tibble_3.3.1 xfun_0.57
#> [58] knitr_1.51 dichromat_2.0-0.1 farver_2.1.2
#> [61] htmltools_0.5.9 rmarkdown_2.31 compiler_4.4.3
#> [64] S7_0.2.2