Functional fine-mapping with fSuSiE
William Denault
Source:vignettes/fsusie_intro.Rmd
fsusie_intro.RmdOverview
fSuSiE extends the SuSiE Bayesian variable selection model to functional traits. The functional response is sampled at multiple positions of a regular grid (e.g., methylation along a CpG island, RNA-seq coverage along a gene body, ATAC-seq across a peak, ChIP-seq read counts on a window). fSuSiE places a wavelet-basis prior on the per-position effect of each variant and performs sparse variable selection by the SuSiE iterative Bayesian stepwise selection (IBSS) algorithm.
The model is
{\bf Y} = {\bf X} {\bf B} + {\bf E},
where {\bf Y} is n x T
(functional response), {\bf X} is
n x p (genotypes), and {\bf
B} is p x T with most rows zero (few causal
variants). fSuSiE writes {\bf B} as a
sum of single-effect matrices {\bf B} =
\sum_{l=1}^L {\bf B}^{(l)} where each {\bf B}^{(l)} has at most one non-zero row,
then fits the model in the wavelet domain.
fsusie() is the single-outcome wrapper around
mfsusie(). The argument order is
(Y, X, pos, ...).
Example data
library(mfsusieR)
data(coverage_example)
str(coverage_example, max.level = 1)
#> List of 6
#> $ X : num [1:574, 1:100] -0.0209 -0.0209 -0.0209 -0.0209 -0.0209 ...
#> $ Y : num [1:574, 1:128] 3.024 0.242 -0.658 0.505 -1.734 ...
#> $ pos : int [1:128] 1 2 3 4 5 6 7 8 9 10 ...
#> $ causal_snps : int [1:2] 25 75
#> $ causal_betas: num [1:2, 1:128] 1.93 -1.36 2.34 -1.31 1.81 ...
#> $ description : chr "Simulated coverage-QTL example. n = 574, p = 100 SNPs from the susieR::N3finemapping cis window, T = 128 evenly"| __truncated__
coverage_example$causal_snps
#> [1] 25 75The data are simulated cis-QTL coverage on a power-of-two grid
(n = 574, p = 100, T = 128). The
genotype matrix is the real LD scaffold from
susieR::N3finemapping$X[, 1:100]. Two causal SNPs at
positions 25 and 75 carry smooth random per-position effects sampled
from the IBSS wavelet prior, the same prior class fSuSiE assumes. The
example is assay-agnostic: the per-position response could be RNA-seq
exon-body coverage, ATAC-seq peak coverage, WGBS / ChIP-seq read counts
on a window, or any per-position coverage / read-count assay.
plot(coverage_example$causal_betas[1L, ], type = "l",
col = "#1f78b4", lwd = 2,
ylim = range(coverage_example$causal_betas),
xlab = "position", ylab = "effect")
abline(h = 0, lty = 3, col = "grey50")
lines(coverage_example$causal_betas[2L, ], col = "#33a02c", lwd = 2)
legend("topright",
legend = paste0("variable ", coverage_example$causal_snps),
col = c("#1f78b4", "#33a02c"), lwd = 2, bty = "n")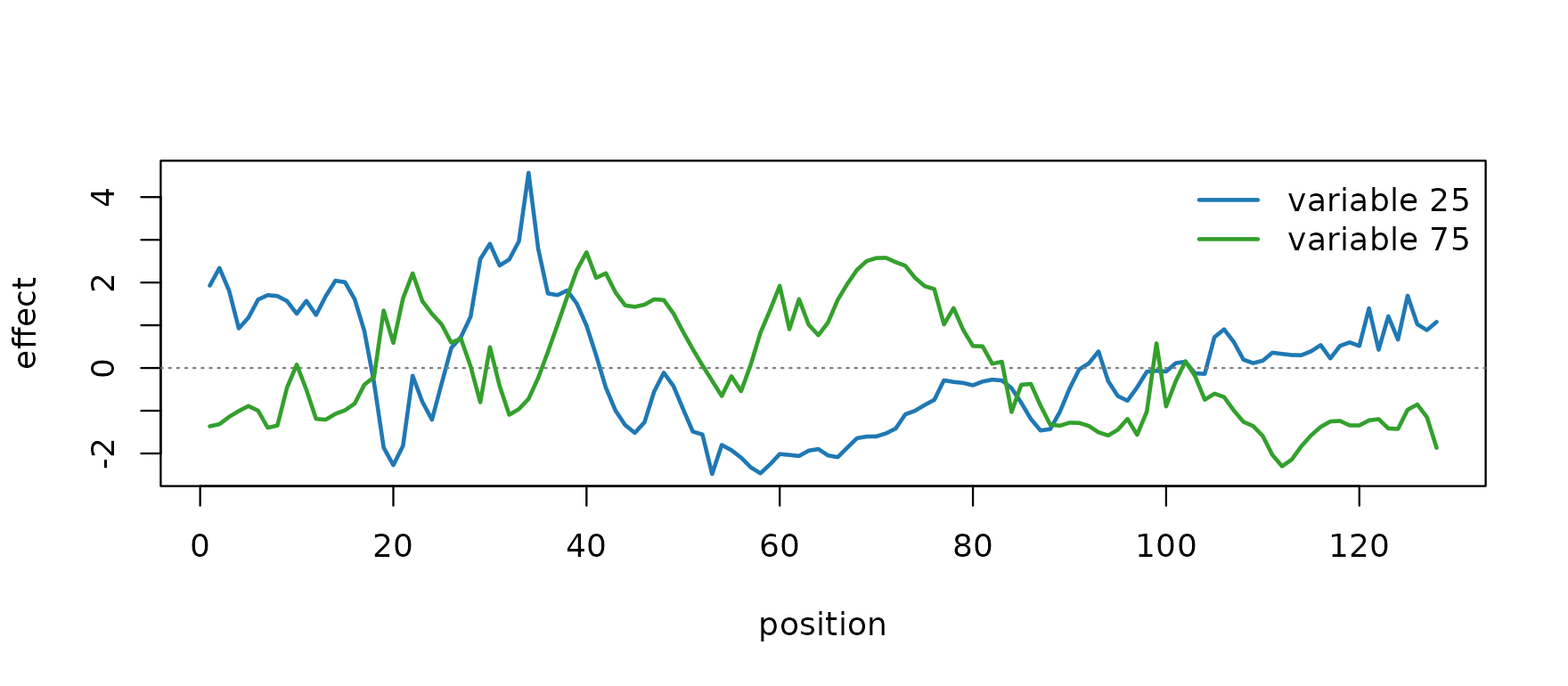
Fitting fSuSiE
fit <- fsusie(coverage_example$Y, coverage_example$X,
pos = coverage_example$pos,
L = 15, L_greedy = 5, verbose = TRUE)
fit$niter; fit$converged
#> [1] 2
#> [1] TRUE
fit$sets$cs
#> $L1
#> [1] 75 83
#>
#> $L2
#> [1] 25
fit$pip[coverage_example$causal_snps]
#> [1] 1.0000000 0.5148505Both causal SNPs receive PIP near 1 and land in two separate credible sets.
Visualizing the fit
mfsusie_plot() returns a two-panel layout for
single-outcome fits: PIPs on top, per-CS effect curves on the bottom.
With facet_cs = "auto" (the default) two CSes whose
affected regions are disjoint are stacked into separate rows.
plot_dims_raw <- mfsusie_plot_dimensions(fit)
plot_dims_raw
#> $width
#> [1] 8
#>
#> $height
#> [1] 6
#>
#> $n_cells
#> [1] 2
mfsusie_plot(fit, effect_variables = coverage_example$causal_snps)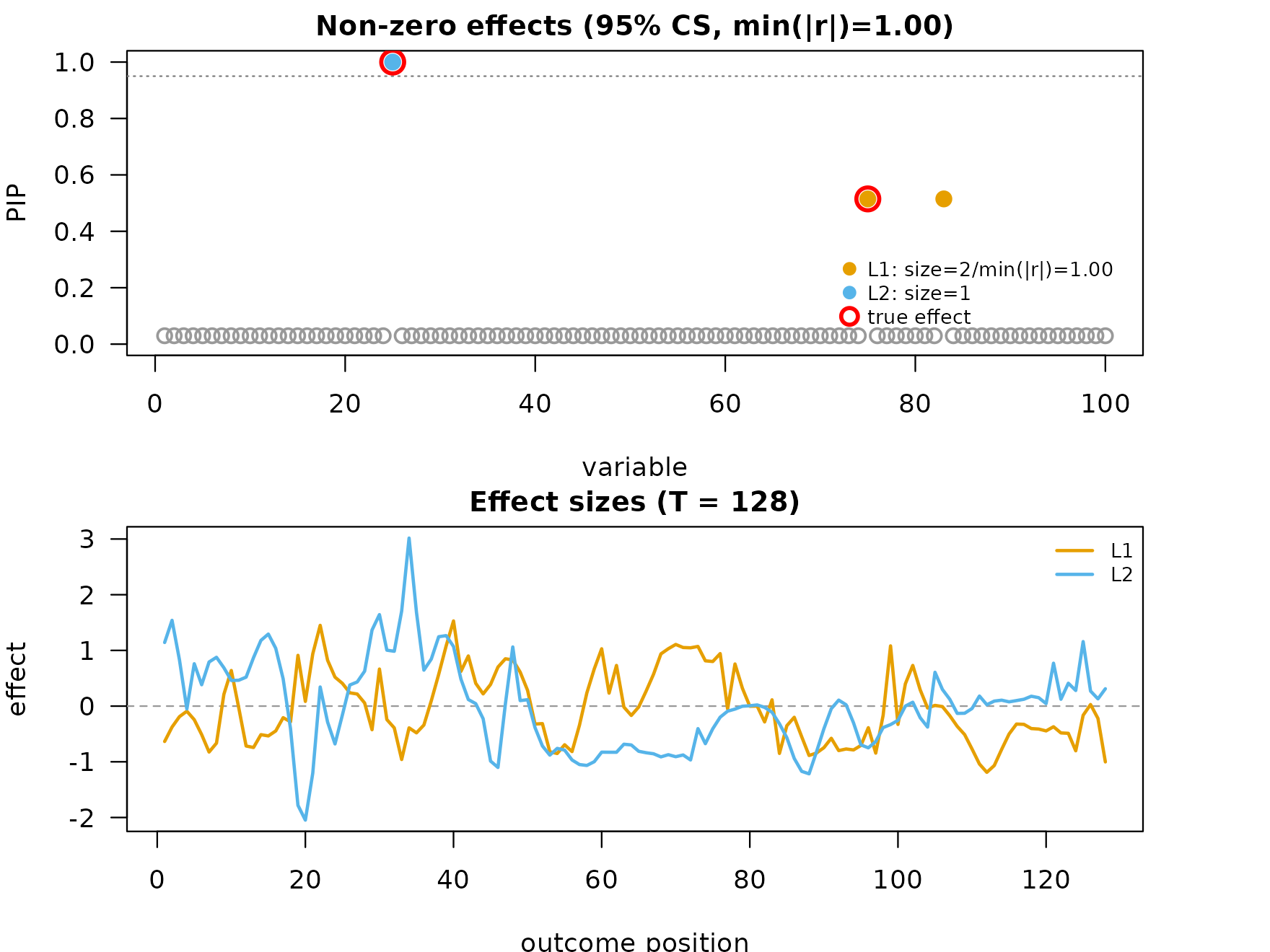
The raw inverse-DWT effect curves track the true shapes but carry
high-frequency content at the smaller wavelet scales. After
post-smoothing with the default method ("TI"), the curves
are denoised and pointwise 95% credible bands are available.
fit_s <- mf_post_smooth(fit)
plot_dims_s <- mfsusie_plot_dimensions(fit_s)
plot_dims_s
#> $width
#> [1] 8
#>
#> $height
#> [1] 6
#>
#> $n_cells
#> [1] 2
mfsusie_plot(fit_s, effect_variables = coverage_example$causal_snps)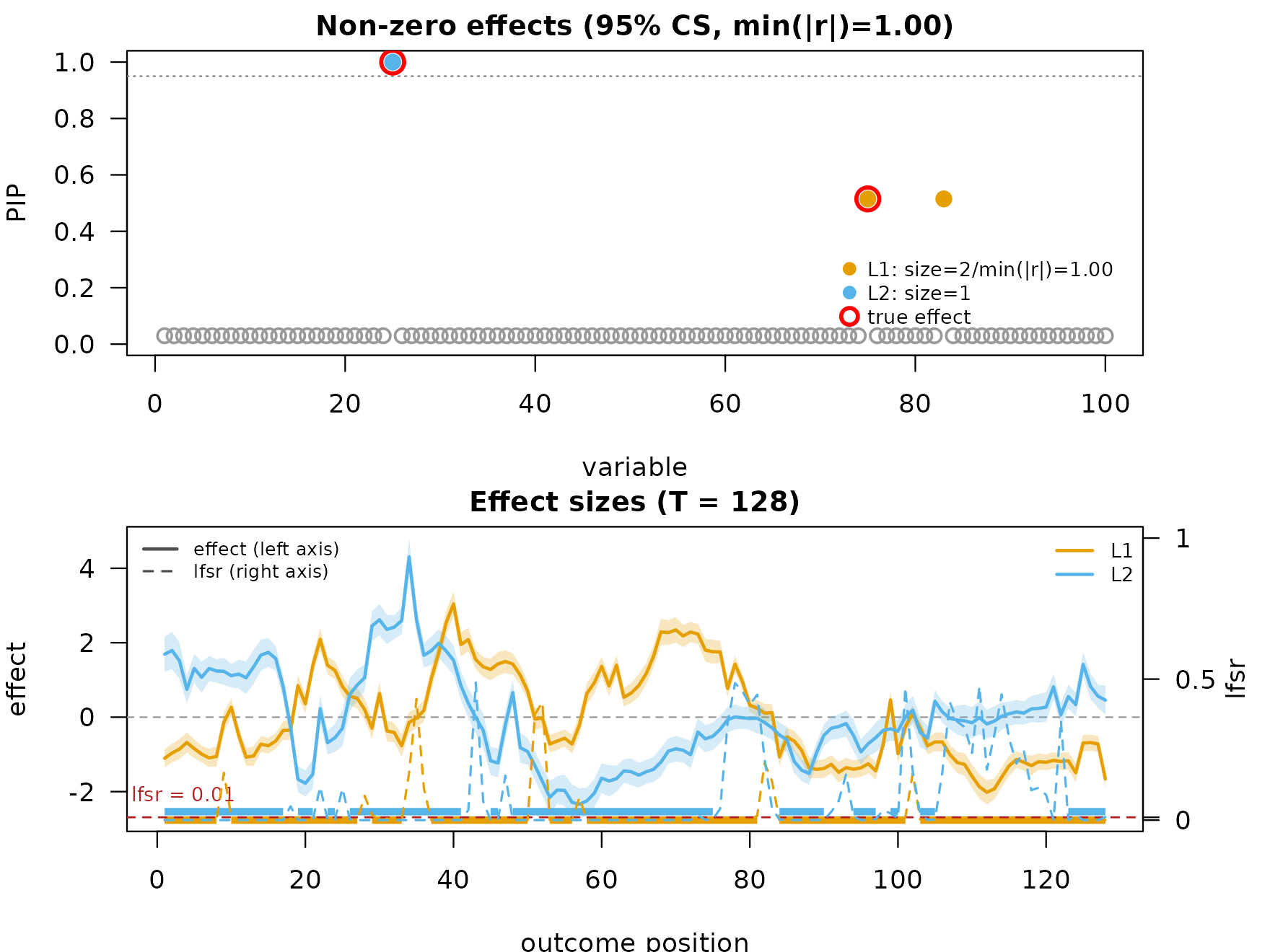
mf_post_smooth() exposes other smoothers
("scalewise", "HMM", "smash") and
their tunable parameters; see vignette("post_processing")
for the comparison.
The errorbar style draws a per-position dot at the posterior mean and a vertical bar to the credible band.
plot_dims_eb <- mfsusie_plot_dimensions(fit_s, facet_cs = "stack")
mfsusie_plot(fit_s, effect_style = "errorbar", facet_cs = "stack",
effect_variables = coverage_example$causal_snps)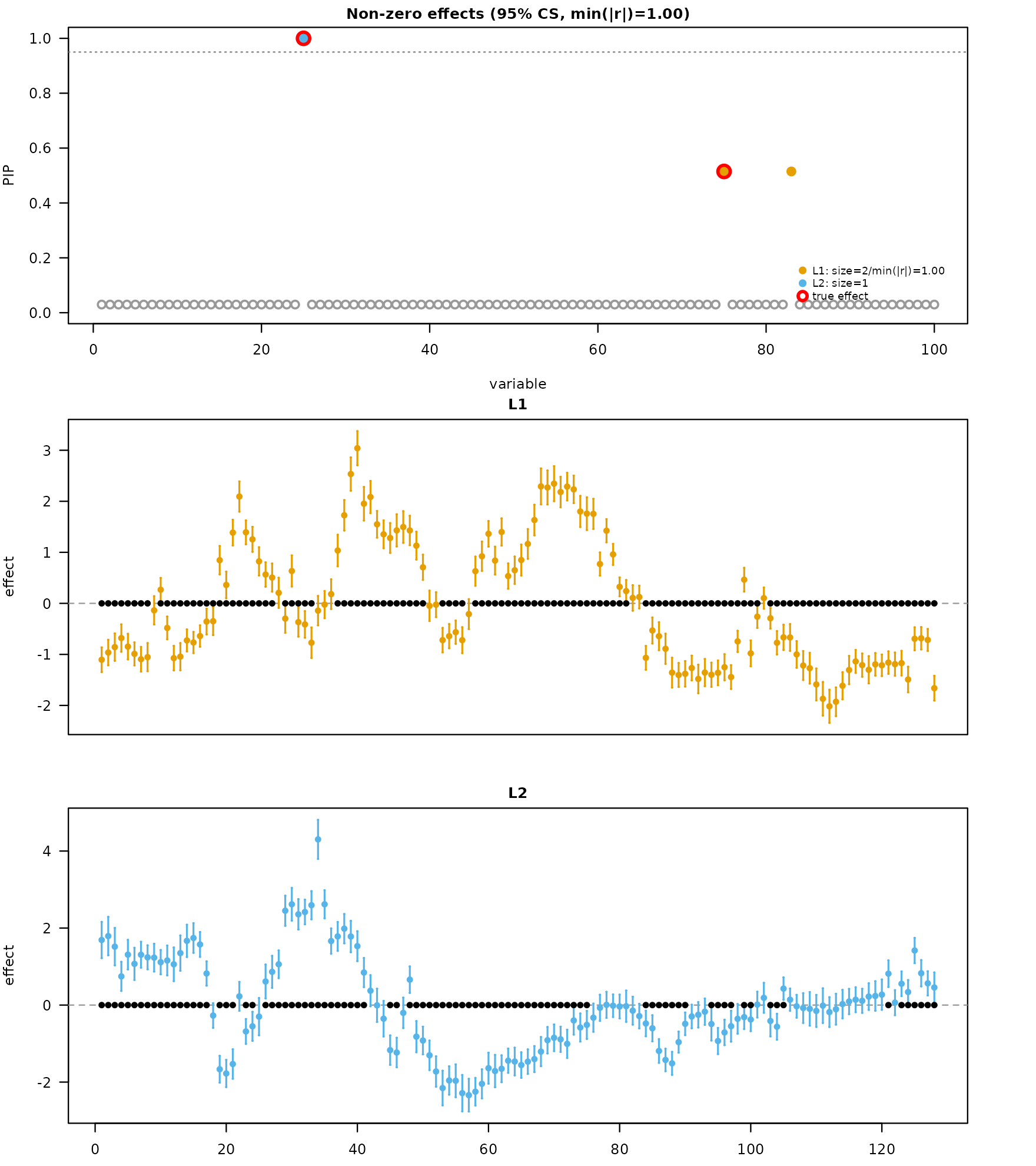
Recovering the true effect
Overlay the TI-smoothed effect against the truth to verify recovery on each causal SNP. The truth is a per-position discrete value, so render it as dots rather than a curve.
truth <- coverage_example$causal_betas
band <- fit_s$smoothed$TI$credible_bands
# Match each fitted CS to its causal SNP by lead PIP.
cs_lead <- vapply(fit$sets$cs, function(idx) {
idx[which.max(fit$pip[idx])]
}, integer(1L))
par(mfrow = c(1L, length(cs_lead)),
mar = c(4, 4, 2.5, 1))
for (i in seq_along(cs_lead)) {
truth_idx <- match(cs_lead[i], coverage_example$causal_snps)
pos_grid <- coverage_example$pos
b_i <- band[[1L]][[i]]
rec_i <- fit_s$smoothed$TI$effect_curves[[1L]][[i]]
yrng <- range(b_i, truth[truth_idx, ], rec_i, na.rm = TRUE)
plot(pos_grid, truth[truth_idx, ], pch = 20L, col = "grey30",
cex = 0.7,
xlab = "position", ylab = "effect",
ylim = yrng,
main = sprintf("CS %d (SNP %d)", i, cs_lead[i]))
polygon(c(pos_grid, rev(pos_grid)),
c(b_i[, 1L], rev(b_i[, 2L])),
col = adjustcolor("#1f78b4", alpha.f = 0.25),
border = NA)
lines(pos_grid, rec_i, col = "#1f78b4", lwd = 2)
abline(h = 0, lty = 3, col = "grey50")
legend("topright", legend = c("truth", "TI smooth", "95% band"),
col = c("grey30", "#1f78b4", adjustcolor("#1f78b4", alpha.f = 0.5)),
lty = c(NA, 1, NA), lwd = c(NA, 2, NA),
pch = c(20, NA, 15), bty = "n", cex = 0.75)
}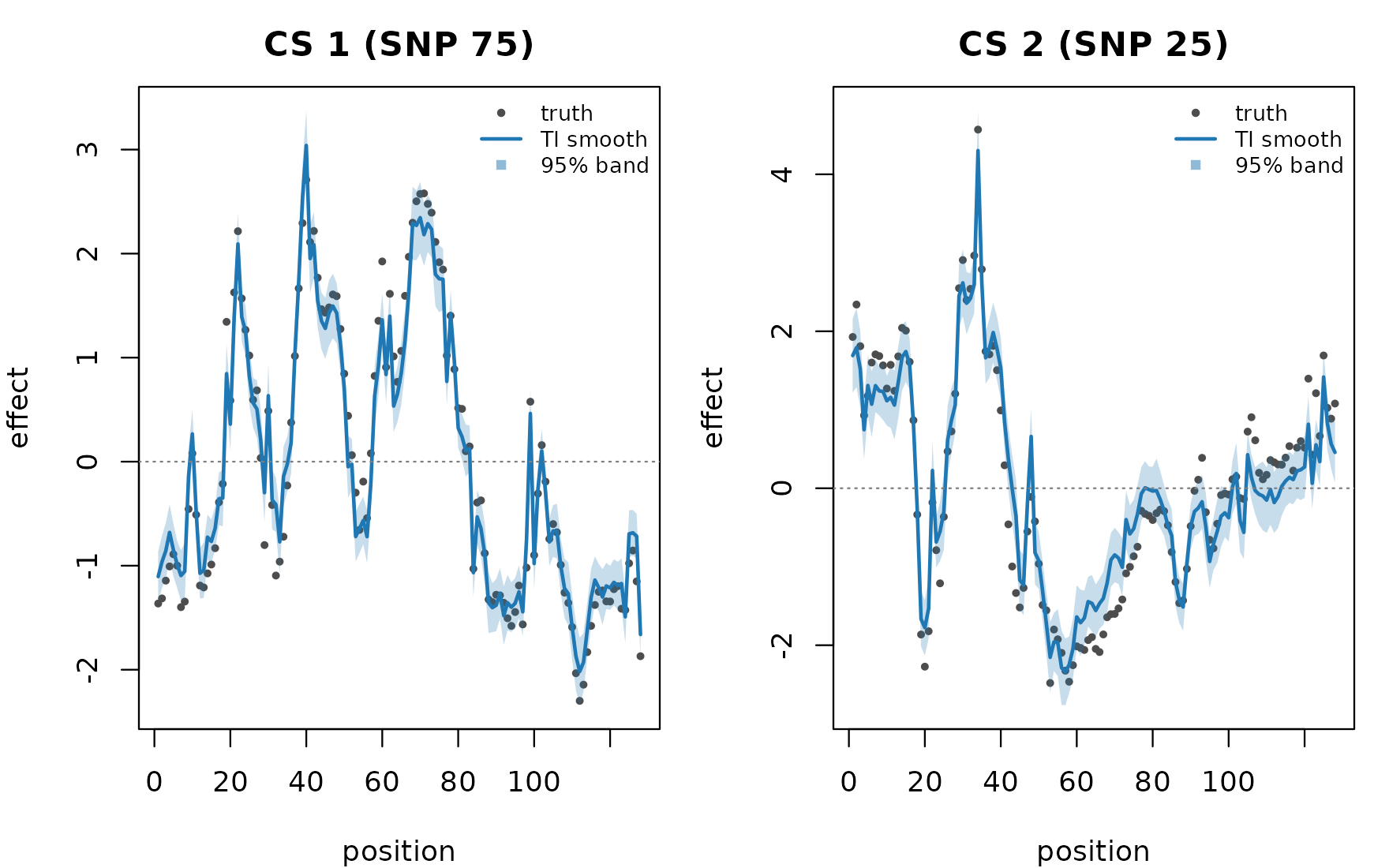
The TI-smoothed curve tracks the truth across the support; the 95% band covers the per-position truth at most sites and only crosses the band on the small high-frequency wiggles the smoother trims.
Uneven sampling positions
Wavelets assume a 2^J evenly-spaced grid.
fsusie() accepts unevenly-spaced positions by remapping the
data onto a regular dyadic grid with the Kovac-Silverman procedure. Pass
the positions through the pos argument; the response is
interpolated internally and the fit returns the per-position effect on
the remapped grid.
To illustrate, drop a random subset of positions from the evenly-spaced response and refit on the irregular grid:
set.seed(2)
T_full <- ncol(coverage_example$Y)
keep <- sort(sample.int(T_full, size = 100L, replace = FALSE))
Y_uneven <- coverage_example$Y[, keep]
fit_uneven <- fsusie(Y_uneven, coverage_example$X,
pos = keep,
L = 15, L_greedy = 5, verbose = TRUE)
fit_uneven$sets$cs
#> $L1
#> [1] 75 83
#>
#> $L2
#> [1] 25
fit_uneven$pip[coverage_example$causal_snps]
#> [1] 1.0000000 0.5148505
fit_uneven_s <- mf_post_smooth(fit_uneven)
plot_dims_u <- mfsusie_plot_dimensions(fit_uneven_s)
mfsusie_plot(fit_uneven_s, effect_variables = coverage_example$causal_snps)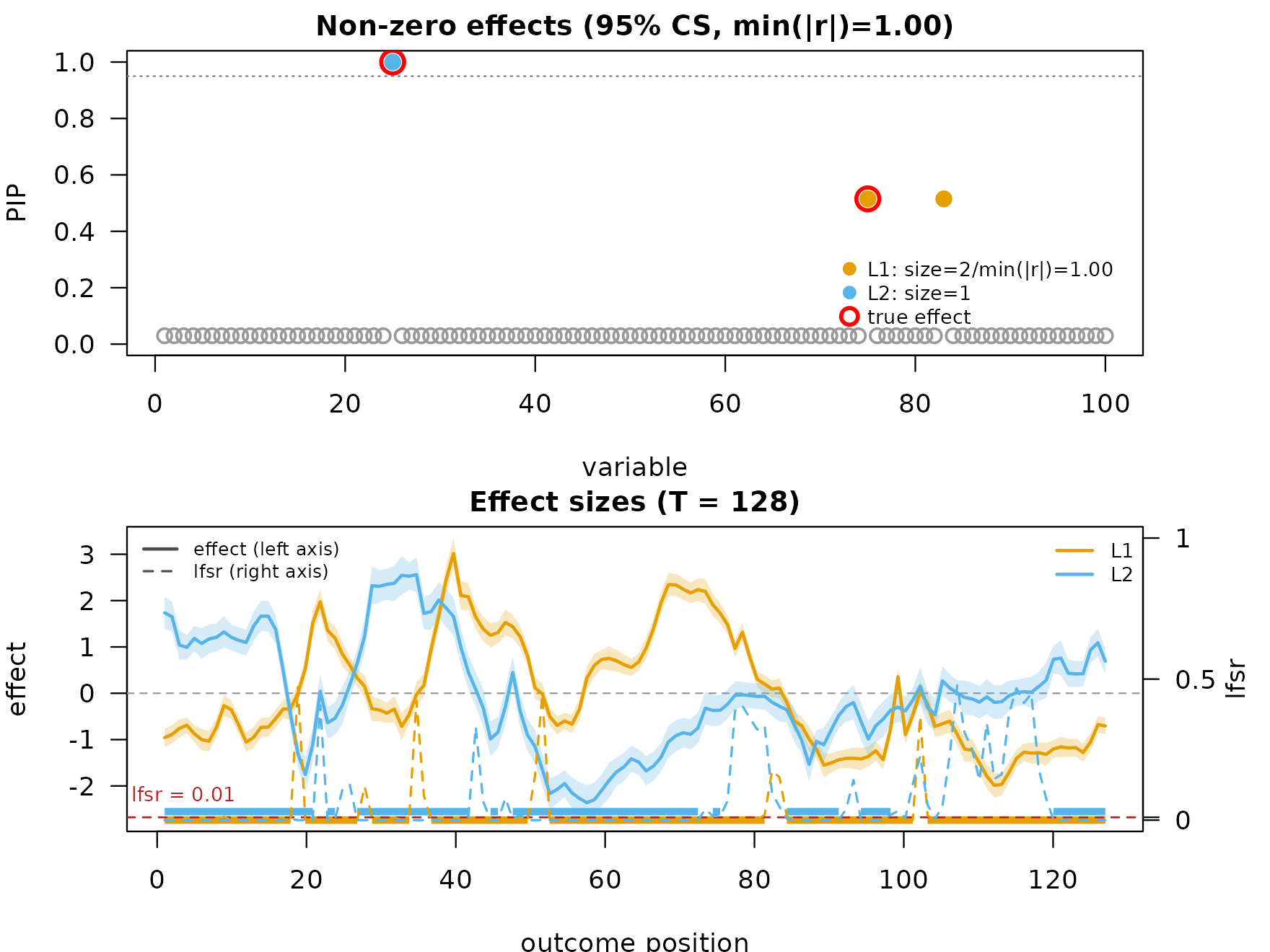
The credible bands are wider than in the evenly-sampled fit because each position now contributes information at a non-uniform local density.
Prior choices
The wavelet-domain mixture prior on {\bf
B}^{(l)} has two options for sharing variance components. The
default, prior_variance_scope = "per_outcome", fits one
mixture weight vector per outcome (collapsed across scales). The
alternative, prior_variance_scope = "per_scale", fits a
separate weight vector at each wavelet scale; this is the original
fSuSiE prior and can yield higher power and estimation accuracy on
signals whose shape varies systematically with scale. Per-scale costs
roughly S_m-fold more per IBSS iter (one
mixsqp call per scale per outcome instead of one per
outcome), so it is opt-in.
fit_per_out <- fsusie(coverage_example$Y, coverage_example$X,
pos = coverage_example$pos,
L = 15, L_greedy = 5,
verbose = FALSE) # per-outcome (default)
fit_per_scale <- fsusie(coverage_example$Y, coverage_example$X,
pos = coverage_example$pos,
L = 15, L_greedy = 5,
prior_variance_scope = "per_scale",
verbose = FALSE)
fit_per_out$niter
#> [1] 2
fit_per_scale$niter
#> [1] 2
plot(fit_per_out$pip, fit_per_scale$pip,
pch = 16, cex = 0.8, col = "#1f78b4",
xlab = "PIP (per-outcome prior, default)",
ylab = "PIP (per-scale prior)",
xlim = c(0, 1), ylim = c(0, 1),
main = "Prior comparison: PIPs")
abline(0, 1, lty = 3, col = "grey50")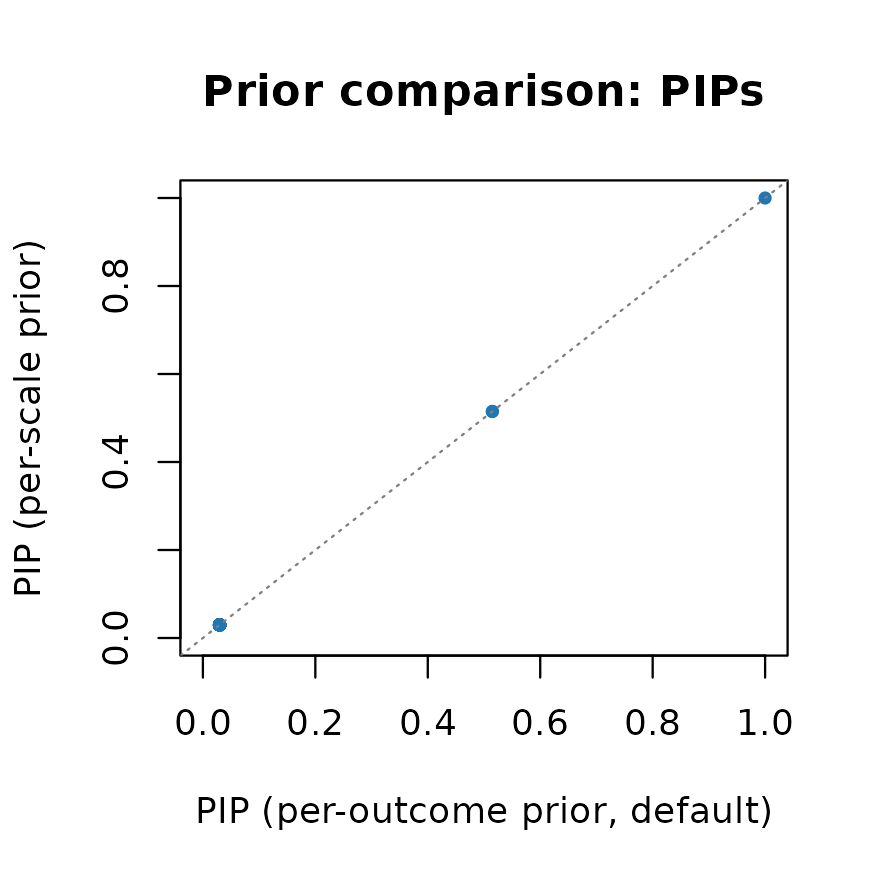
The PIPs are nearly identical for the variants in either credible set. The per-scale prior is the recommended default for functional fine-mapping problems.
Wavelet-domain preprocessing
fsusie() exposes two preprocessing flags for the wavelet
matrix:
-
wavelet_magnitude_cutoff(default0): wavelet columns withmedian(|column|) <= wavelet_magnitude_cutoffare masked, contributing(Bhat = 0, Shat = 1)to the SER step. Use a small positive value when the response has sparse non-zero entries (e.g., ATAC-seq read pileups). -
wavelet_qnorm(defaultTRUE): applies a column-wise rank-based normal quantile transform to the wavelet matrix. Useful when the per-position residuals are heavy-tailed or visibly non-Gaussian. Set toFALSEto skip the transform when the wavelet coefficients are already approximately Gaussian.
See ?mfsusie for the full set of preprocessing
options.
What the fit object carries
str(fit, max.level = 1)
#> List of 31
#> $ alpha : num [1:5, 1:100] 0.00 1.03e-284 1.00e-02 1.00e-02 1.00e-02 ...
#> $ mu :List of 5
#> $ mu2 :List of 5
#> $ V_grid :List of 1
#> $ pi_V :List of 1
#> $ null_prior_weight : num 2
#> $ G_prior :List of 1
#> $ cross_outcome_combiner: list()
#> ..- attr(*, "class")= chr [1:2] "mf_prior_cross_outcome_independent" "mf_prior_cross_outcome"
#> $ KL : num [1:5] 1.38e+02 1.01e+02 -1.49e-06 -1.49e-06 -1.49e-06
#> $ lbf : num [1:5] 9.35e+02 6.29e+02 1.49e-06 1.49e-06 1.49e-06
#> $ lbf_variable : num [1:5, 1:100] -22.8 -19.8 0 0 0 ...
#> $ lbf_variable_outcome : num [1:5, 1:100, 1] -22.8 -19.8 0 0 0 ...
#> $ sigma2 :List of 1
#> $ pi : num [1:100] 0.01 0.01 0.01 0.01 0.01 0.01 0.01 0.01 0.01 0.01 ...
#> $ fitted :List of 1
#> $ intercept :List of 1
#> $ L : int 5
#> $ p : int 100
#> $ M : int 1
#> $ null_index : num 0
#> $ converged : logi TRUE
#> $ em_cache :List of 4
#> $ ser_cache :List of 3
#> $ elbo : num [1:2] NA -102703
#> $ niter : int 2
#> $ X_column_scale_factors: num [1:100] 0.143 0.624 0.684 0.23 0.411 ...
#> $ sets :List of 5
#> $ pip : num [1:100] 0.0297 0.0297 0.0297 0.0297 0.0297 ...
#> $ Y_grid :List of 1
#> $ X_eff :List of 5
#> $ dwt_meta :List of 12
#> - attr(*, "class")= chr [1:2] "mfsusie" "susie"The fit is a list of class c("mfsusie", "susie") with
the standard SuSiE slots (alpha, mu,
mu2, lbf, lbf_variable,
KL, pip, sets) plus per-outcome
wavelet metadata (dwt_meta) used by
predict.mfsusie, coef.mfsusie, and
fitted.mfsusie. After
mf_post_smooth(fit, method = ...), the chosen smoother’s
output lives at fit$smoothed[[method]] (with
effect_curves, credible_bands, and an optional
lfsr_curves slot for HMM). The fit accumulates per-method
results across calls, so multiple smoothers can coexist on the same fit;
see vignette("post_processing") for the comparison
workflow. mfsusie_plot() picks a smoother by priority
TI > smash > HMM > scalewise when more than one is
present.
Session info
This is the version of R and the packages that were used to generate these results.
sessionInfo()
#> R version 4.4.3 (2025-02-28)
#> Platform: x86_64-conda-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS/LAPACK: /home/runner/work/mfsusieR/mfsusieR/.pixi/envs/r44/lib/libopenblasp-r0.3.32.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: Etc/UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] mfsusieR_0.0.2
#>
#> loaded via a namespace (and not attached):
#> [1] sass_0.4.10 generics_0.1.4 ashr_2.2-63
#> [4] lattice_0.22-9 magrittr_2.0.5 digest_0.6.39
#> [7] evaluate_1.0.5 grid_4.4.3 RColorBrewer_1.1-3
#> [10] fastmap_1.2.0 plyr_1.8.9 jsonlite_2.0.0
#> [13] Matrix_1.7-5 reshape_0.8.10 mixsqp_0.3-54
#> [16] scales_1.4.0 truncnorm_1.0-9 invgamma_1.2
#> [19] textshaping_1.0.5 jquerylib_0.1.4 cli_3.6.6
#> [22] rlang_1.2.0 zigg_0.0.2 crayon_1.5.3
#> [25] LaplacesDemon_16.1.8 susieR_0.16.1 cachem_1.1.0
#> [28] yaml_2.3.12 otel_0.2.0 tools_4.4.3
#> [31] SQUAREM_2026.1 parallel_4.4.3 dplyr_1.2.1
#> [34] wavethresh_4.7.3 ggplot2_4.0.3 Rfast_2.1.5.2
#> [37] vctrs_0.7.3 R6_2.6.1 matrixStats_1.5.0
#> [40] lifecycle_1.0.5 fs_2.1.0 htmlwidgets_1.6.4
#> [43] MASS_7.3-65 ragg_1.5.2 irlba_2.3.7
#> [46] pkgconfig_2.0.3 desc_1.4.3 pillar_1.11.1
#> [49] pkgdown_2.2.0 RcppParallel_5.1.11-2 bslib_0.10.0
#> [52] gtable_0.3.6 glue_1.8.1 Rcpp_1.1.1-1.1
#> [55] systemfonts_1.3.2 tidyselect_1.2.1 tibble_3.3.1
#> [58] xfun_0.57 knitr_1.51 dichromat_2.0-0.1
#> [61] farver_2.1.2 htmltools_0.5.9 rmarkdown_2.31
#> [64] compiler_4.4.3 S7_0.2.2