Cis-QTL Analysis and GWAS Integration Workflow
Anqi Wang and Gao Wang
2026-05-29
Source:vignettes/cis-analysis.Rmd
cis-analysis.RmdHere we demonstrate the use of the pecotmr package to
perform colocalization, TWAS and enrichment analysis between a pair of
molecular QTL and GWAS using de-identified example data shipped with the
package. All sample names, variant positions, and identifiers are
synthetic.
Load data
data(gwas_sumstats_example)
data(eqtl_region_example)
gwas <- gwas_sumstats_example
eqtl <- eqtl_region_example
head(gwas)## variant_id chrom pos A1 A2 beta se z
## 1 chr22:32119788:T:C chr22 32119788 T C -0.0089 0.0111 -0.8018018
## 2 chr22:32119867:T:G chr22 32119867 T G -0.0089 0.0111 -0.8018018
## 3 chr22:32119961:T:G chr22 32119961 T G 0.0236 0.0083 2.8433735
## 4 chr22:32120053:T:C chr22 32120053 T C -0.0217 0.0246 -0.8821138
## 5 chr22:32120593:A:G chr22 32120593 A G -0.0232 0.0246 -0.9430894
## 6 chr22:32120636:T:C chr22 32120636 T C 0.0045 0.0141 0.3191489
names(eqtl)## [1] "X" "y_res"
head(eqtl$X[, 1:5])## chr22:32119788:T:C chr22:32119867:T:G chr22:32119961:T:G
## sample_001 0 0 0
## sample_002 0 0 1
## sample_003 1 1 0
## sample_004 1 1 1
## sample_005 1 1 1
## sample_006 1 1 1
## chr22:32120053:T:C chr22:32120593:A:G
## sample_001 0 0
## sample_002 0 0
## sample_003 0 0
## sample_004 0 0
## sample_005 0 0
## sample_006 0 0
head(eqtl$y_res)## sample_001 sample_002 sample_003 sample_004 sample_005 sample_006
## 0.50197972 0.24567929 -0.06281688 -0.53649517 -0.56143407 -0.54624170
dim(eqtl$X)## [1] 415 2828Fine-mapping using SuSiE
## ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
## ✔ dplyr 1.2.1 ✔ readr 2.2.0
## ✔ forcats 1.0.1 ✔ stringr 1.6.0
## ✔ ggplot2 4.0.3 ✔ tibble 3.3.1
## ✔ lubridate 1.9.5 ✔ tidyr 1.3.2
## ✔ purrr 1.2.2
## ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
## ✖ dplyr::filter() masks stats::filter()
## ✖ dplyr::lag() masks stats::lag()
## ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
get_cs_index <- function(snps_idx, fitted_data) {
idx <- tryCatch(
which(
pmap(list(a = fitted_data$sets$cs), function(a) snps_idx %in% a) %>% unlist()
),
error = function(e) NA_integer_
)
if(length(idx) == 0) return(NA_integer_)
return(idx)
}
signal_cutoff = 0.1
coverage = 0.7
susie_res <- susie(eqtl$X, eqtl$y_res,
L=20,
max_iter=500,
estimate_residual_variance=TRUE,
estimate_prior_variance=TRUE,
refine=TRUE,
compute_univariate_zscore=FALSE,
coverage=coverage)
secondary_coverage = 0.2
susie_res$sets_secondary = susie_get_cs(susie_res, eqtl$X, coverage=secondary_coverage)
compute_maf <- function(geno){
f <- mean(geno,na.rm = TRUE)/2
return(min(f, 1-f))
}
variants_index = c(which(susie_res$pip >= signal_cutoff), unlist(susie_res$sets$cs)) %>% unique %>% sort
maf = apply(eqtl$X, 2, compute_maf)[variants_index]
all_variants = colnames(eqtl$X)
variants = all_variants[variants_index]
pip = susie_res$pip[variants_index]
cs_info = map_int(variants_index, ~get_cs_index(.x, susie_res))
cs_index = ifelse(is.na(cs_info), 0, str_replace(names(susie_res$sets$cs)[cs_info], "L", "") %>% as.numeric)
univariate_res = univariate_regression(eqtl$X[, variants_index, drop=F], eqtl$y_res)
# here betahat and sebetahat were computed from standardized X and Y
susie_res$top_loci = cbind(variants, maf, univariate_res$betahat, univariate_res$sebetahat, pip, cs_index)
colnames(susie_res$top_loci) = c("variant_id", "maf", "bhat", "sbhat", "pip", "cs_index_primary")
rownames(susie_res$top_loci) = NULL
susie_res$top_loci## variant_id maf bhat
## [1,] "chr22:32596246:T:C" "0.278313253012048" "-0.122514739342209"
## [2,] "chr22:32724254:T:C" "0.472289156626506" "0.146350893456887"
## [3,] "chr22:32724508:C:A" "0.472289156626506" "0.146350893456887"
## [4,] "chr22:32728910:G:A" "0.469879518072289" "0.146101325781137"
## [5,] "chr22:32729588:T:C" "0.471084337349398" "0.145082188952344"
## [6,] "chr22:32732510:G:A" "0.472289156626506" "0.147441999306526"
## [7,] "chr22:32734332:A:G" "0.497590361445783" "0.14373324203229"
## [8,] "chr22:32737583:C:T" "0.496385542168675" "0.142163759153291"
## [9,] "chr22:32737848:A:G" "0.497590361445783" "0.14373324203229"
## [10,] "chr22:32740783:T:C" "0.497590361445783" "0.14373324203229"
## [11,] "chr22:32743285:A:C" "0.497590361445783" "0.14373324203229"
## [12,] "chr22:32745710:T:C" "0.473493975903614" "0.148379062725204"
## [13,] "chr22:32745749:T:C" "0.497590361445783" "0.141364354229342"
## [14,] "chr22:32752021:A:G" "0.49993728060858" "0.141437506070959"
## [15,] "chr22:32755495:C:T" "0.474698795180723" "0.144921704020173"
## sbhat pip cs_index_primary
## [1,] "0.0249868671221125" "0.144496972379586" "0"
## [2,] "0.0235695850612729" "0.0442663476866332" "1"
## [3,] "0.0235695850612729" "0.0442663476866332" "1"
## [4,] "0.0235890553342029" "0.0394962530308444" "1"
## [5,] "0.0235285991106962" "0.0345101548349765" "1"
## [6,] "0.0234182228577676" "0.0684513653202697" "1"
## [7,] "0.0230653145891427" "0.066349050628123" "1"
## [8,] "0.0231535224323869" "0.0411110619568467" "1"
## [9,] "0.0230653145891427" "0.066349050628123" "1"
## [10,] "0.0230653145891427" "0.066349050628123" "1"
## [11,] "0.0230653145891427" "0.066349050628123" "1"
## [12,] "0.0234638029050504" "0.0769631008098507" "1"
## [13,] "0.0231007359525088" "0.0363654786045676" "1"
## [14,] "0.0230989526643856" "0.0363658592244495" "1"
## [15,] "0.0234421257080182" "0.0365496709704958" "1"Top PIP variants:
sort(susie_res$pip, decreasing = TRUE)[1:5]## chr22:32596246:T:C chr22:32745710:T:C chr22:32732510:G:A chr22:32734332:A:G
## 0.14449697 0.07696310 0.06845137 0.06634905
## chr22:32737848:A:G
## 0.06634905TWAS
Use SuSiE weights,
R = cor(eqtl$X)
weights = susie_weights(susie_fit = susie_res)
twas_z(weights, gwas$z, R=R)## $z
## [,1]
## [1,] -3.07806
##
## $pval
## [,1]
## [1,] 0.002083527## [,1]
## [1,] 0.1220979Now select only the top loci,
plot(weights)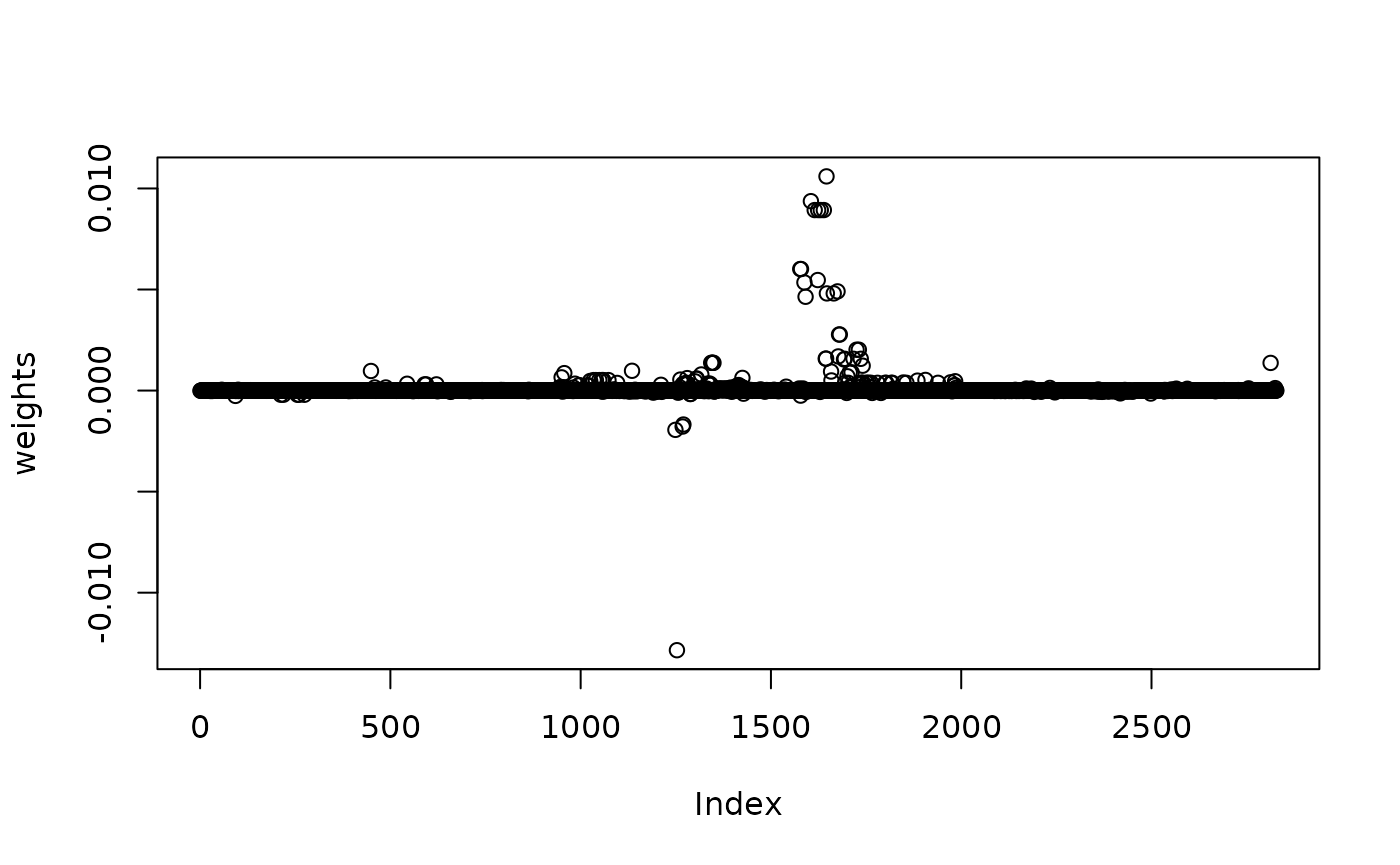
weights[!all_variants %in% susie_res$top_loci[,1]] = 0
plot(weights)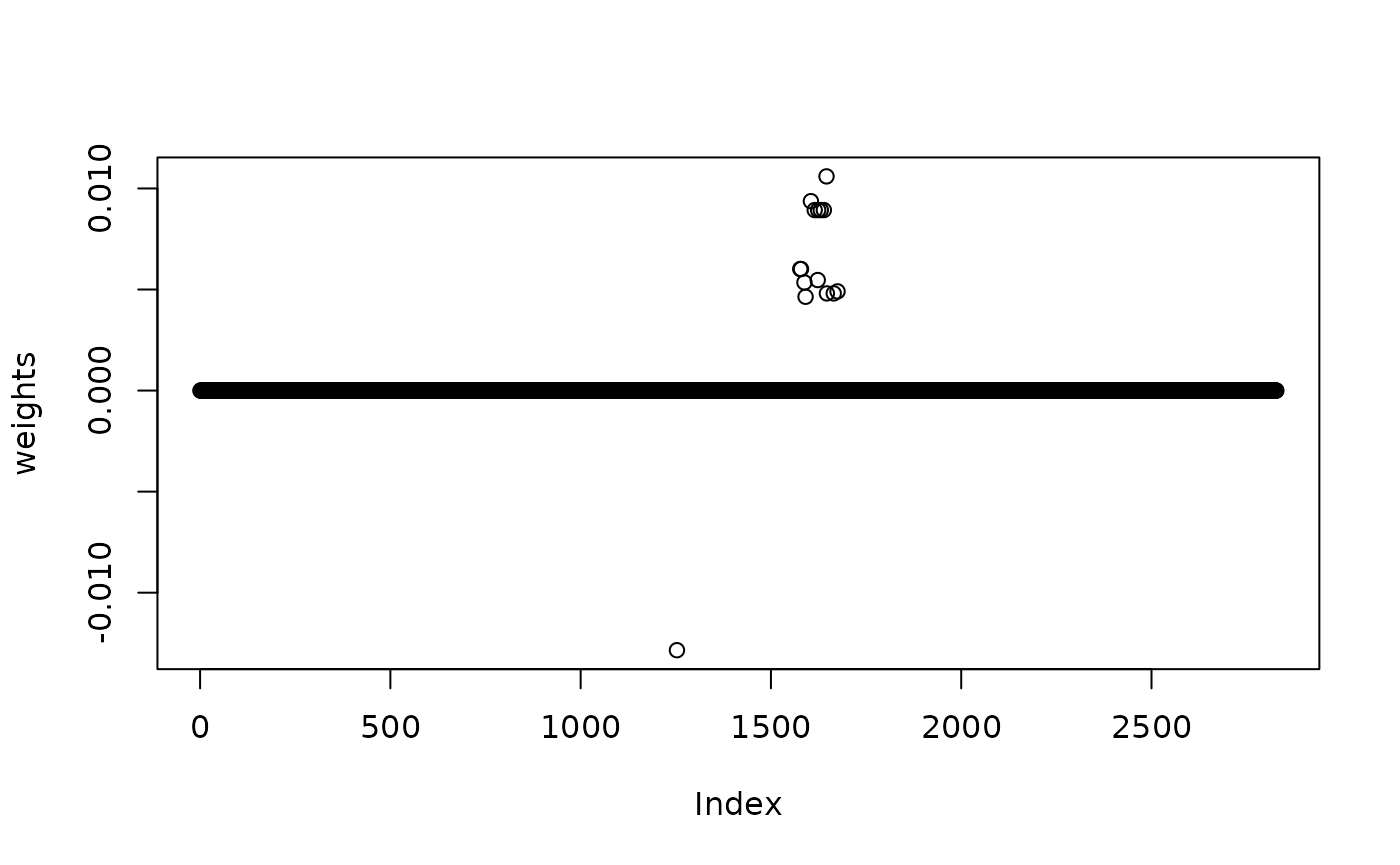
twas_z(weights, gwas$z, R=R)## $z
## [,1]
## [1,] -2.438249
##
## $pval
## [,1]
## [1,] 0.01475859Use elastic net (alpha=0.5),
## [,1]
## [1,] 0.1817924
twas_z(weights, gwas$z, R=R)## $z
## [,1]
## [1,] -4.556306
##
## $pval
## [,1]
## [1,] 5.206117e-06
plot(weights)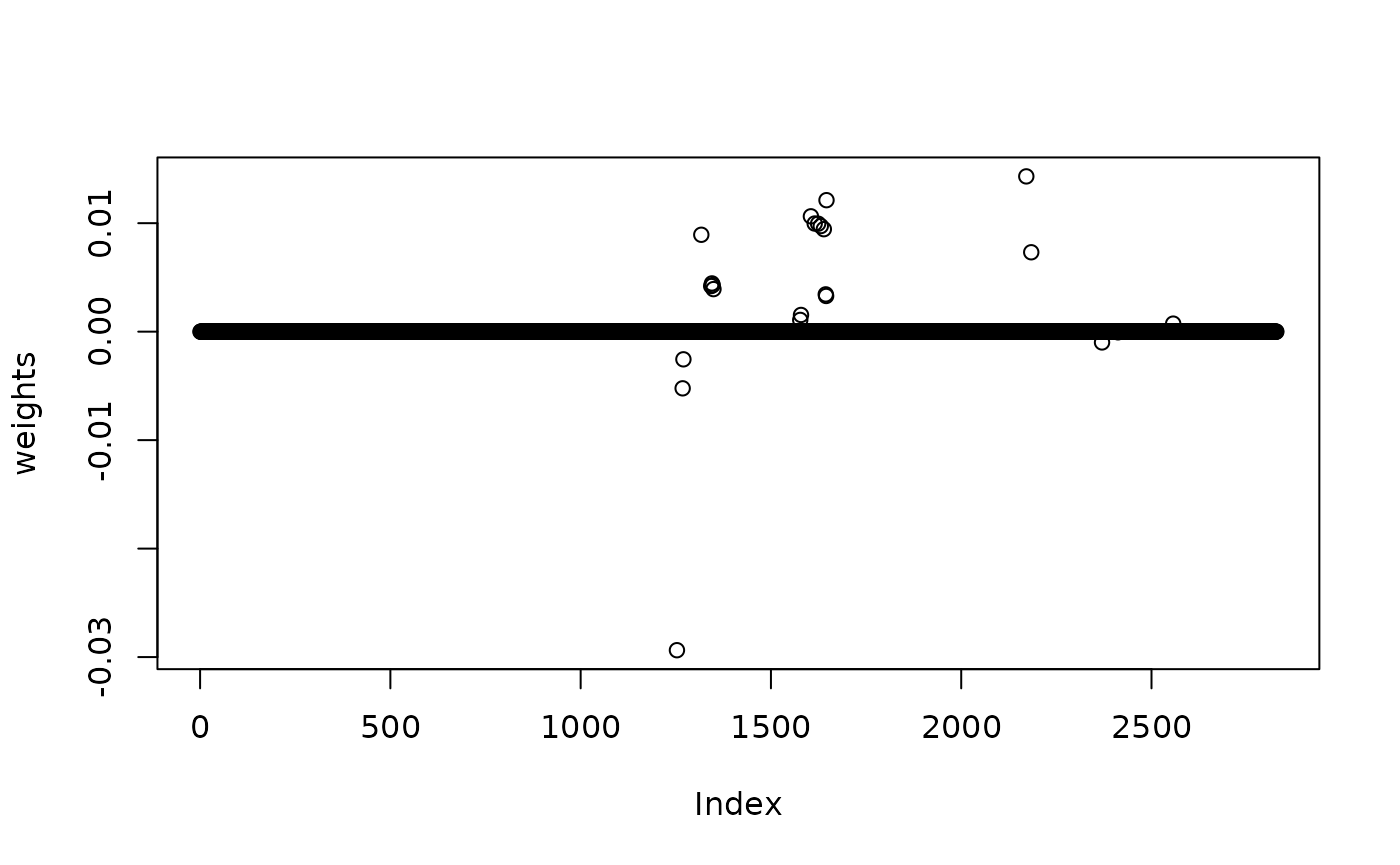
Use LASSO,
## [,1]
## [1,] 0.23199
twas_z(weights, gwas$z, R=R)## $z
## [,1]
## [1,] -4.484433
##
## $pval
## [,1]
## [1,] 7.31079e-06
plot(weights)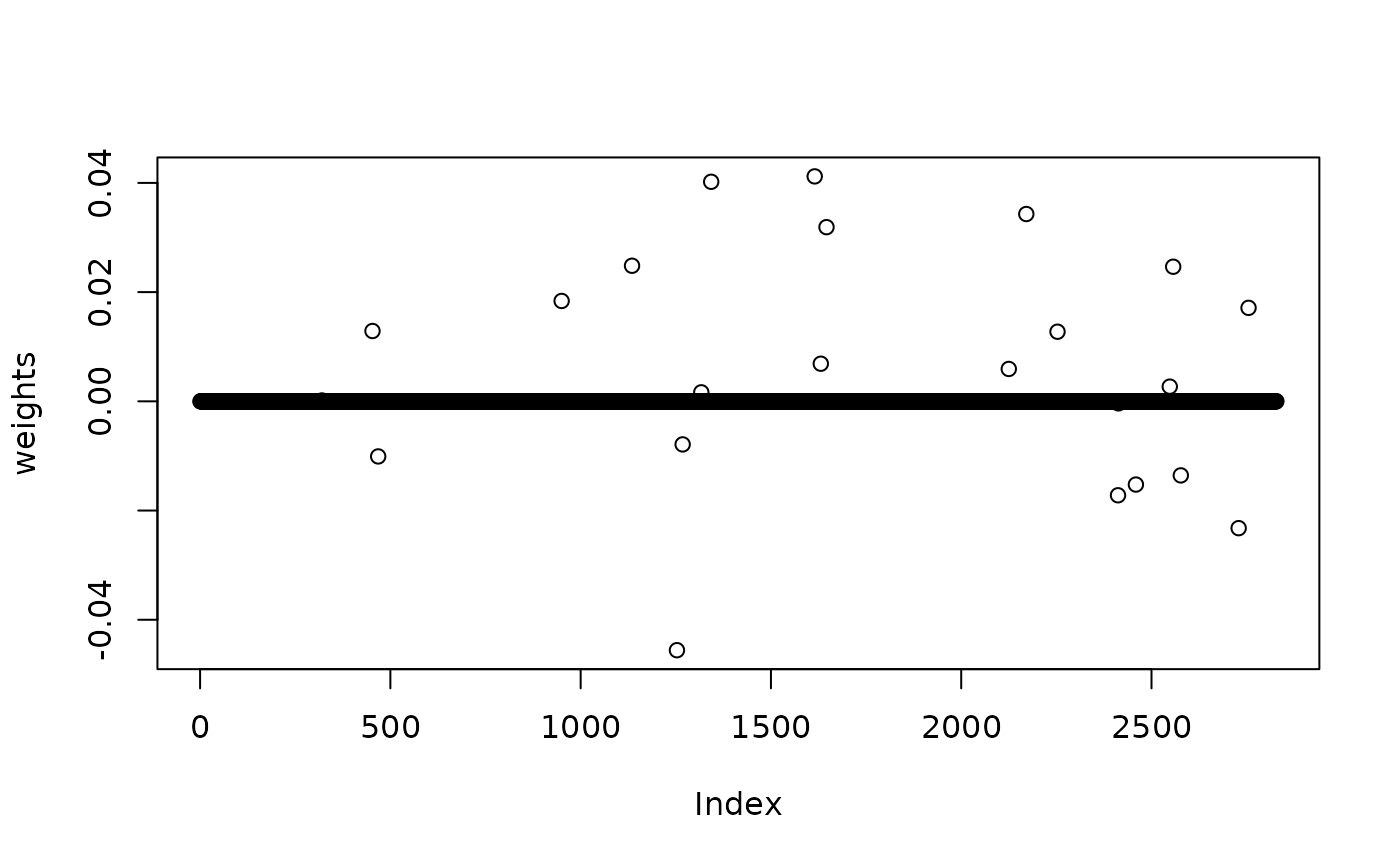
Use mr.ash,
lasso.weights = glmnet_weights(eqtl$X, eqtl$y_res, alpha=1)
weights = mrash_weights(eqtl$X, eqtl$y_res, init_prior_sd=TRUE, beta.init=lasso.weights)## Mr.ASH terminated at iteration 64: max|beta|=2.8637e-03, sigma2=1.0243e-01, pi0=0.9529## [,1]
## [1,] 0.1872491
twas_z(weights, gwas$z, R=R)## $z
## [,1]
## [1,] -3.188965
##
## $pval
## [,1]
## [1,] 0.00142783
plot(weights)
weights = mrash_weights(eqtl$X, eqtl$y_res, init_prior_sd=FALSE, beta.init=lasso.weights)## Mr.ASH terminated at iteration 49: max|beta|=4.5258e-03, sigma2=1.0294e-01, pi0=0.9752## [,1]
## [1,] 0.185199
twas_z(weights, gwas$z, R=R)## $z
## [,1]
## [1,] -3.299771
##
## $pval
## [,1]
## [1,] 0.0009676385
plot(weights)
Enrichment analysis
data(gwas_finemapping_example)
data(qtl_finemapping_example)
gwas_path <- tempfile(fileext = ".rds")
qtl_path <- tempfile(fileext = ".rds")
saveRDS(gwas_finemapping_example, gwas_path)
saveRDS(qtl_finemapping_example, qtl_path)
xqtl_enrichment_wrapper(gwas_files = gwas_path, xqtl_files = qtl_path,
xqtl_finemapping_obj = c("context_1", "susie_result_trimmed"),
xqtl_varname_obj = c("context_1", "variant_names"))## Warning in compute_qtl_enrichment(gwas_pip = dat$gwas_pip, susie_qtl_regions =
## dat$xqtl_data, : num_gwas is not provided. Estimating pi_gwas from the data.
## Note that this estimate may be biased if the input gwas_pip does not contain
## genome-wide variants.## Estimated pi_gwas: 0.00141## Warning in compute_qtl_enrichment(gwas_pip = dat$gwas_pip, susie_qtl_regions =
## dat$xqtl_data, : pi_qtl is not provided. Estimating pi_qtl from the data. Note
## that this estimate may be biased if either 1) the input susie_qtl_regions does
## not have enough data, or 2) the single effects only include variables inside of
## credible sets or signal clusters.## Estimated pi_qtl: 0.00071## Fine-mapped GWAS and QTL data loaded successfully for enrichment analysis!
## Proportion of xQTL missing from GWAS variants: 0 in MI round 0
## Proportion of xQTL missing from GWAS variants: 0 in MI round 1
## Proportion of xQTL missing from GWAS variants: 0 in MI round 2
## Proportion of xQTL missing from GWAS variants: 0 in MI round 3
## Proportion of xQTL missing from GWAS variants: 0 in MI round 4
## Proportion of xQTL missing from GWAS variants: 0 in MI round 5
## Proportion of xQTL missing from GWAS variants: 0 in MI round 6
## Proportion of xQTL missing from GWAS variants: 0 in MI round 7
## Proportion of xQTL missing from GWAS variants: 0 in MI round 8
## Proportion of xQTL missing from GWAS variants: 0 in MI round 9
## Proportion of xQTL missing from GWAS variants: 0 in MI round 10
## Proportion of xQTL missing from GWAS variants: 0 in MI round 11
## Proportion of xQTL missing from GWAS variants: 0 in MI round 12
## Proportion of xQTL missing from GWAS variants: 0 in MI round 13
## Proportion of xQTL missing from GWAS variants: 0 in MI round 14
## Proportion of xQTL missing from GWAS variants: 0 in MI round 15
## Proportion of xQTL missing from GWAS variants: 0 in MI round 16
## Proportion of xQTL missing from GWAS variants: 0 in MI round 17
## Proportion of xQTL missing from GWAS variants: 0 in MI round 18
## Proportion of xQTL missing from GWAS variants: 0 in MI round 19
## Proportion of xQTL missing from GWAS variants: 0 in MI round 20
## Proportion of xQTL missing from GWAS variants: 0 in MI round 21
## Proportion of xQTL missing from GWAS variants: 0 in MI round 22
## Proportion of xQTL missing from GWAS variants: 0 in MI round 23
## Proportion of xQTL missing from GWAS variants: 0 in MI round 24
## EM updates completed!
## Outlier filtering removed 1 MI round(s)## [[1]]
## [[1]]$`Alternative (coloc) p1`
## [1] 0.001413425
##
## [[1]]$`Alternative (coloc) p12`
## [1] 1.000238e-06
##
## [[1]]$`Alternative (coloc) p2`
## [1] 0.0007061743
##
## [[1]]$`Effective MI rounds`
## [1] 24
##
## [[1]]$`Enrichment (no shrinkage)`
## [1] -7.950622
##
## [[1]]$`Enrichment (w/ shrinkage)`
## [1] -7.952562e-06
##
## [[1]]$Intercept
## [1] -6.559616
##
## [[1]]$`sd (intercept)`
## [1] 0.5001639
##
## [[1]]$`sd (no shrinkage)`
## [1] 999.8775
##
## [[1]]$`sd (w/ shrinkage)`
## [1] 0.9999995
##
##
## $unused_xqtl_variants
## $unused_xqtl_variants[[1]]
## NULL