Simulated cis-eQTL data inspired by published GTEx case
studies (AHCYL1, SCD, HSP90AA1, STMN2, ABHD17A): a
functional response on 256 gene-body positions, three
causal SNPs with localized peak effects at distinct 5',
mid-body, and 3' positions. Used by
vignette("fsusie_gtex_case_study").
Format
A list with components
Xn x pgenotype matrix (p = 200) sliced fromsusieR::N3finemapping$X.Yn x Tlog1p-coverage matrix (T = 256).poslength-
Tinteger vector of gene-body positions.causal_snpsinteger vector of column indices in
Xthat carry true effects.causal_betaslength(causal_snps) x Tmatrix of true per-position effects.descriptionfree-text description.
Examples
# \donttest{
data(gtex_example)
fit <- fsusie(gtex_example$Y, gtex_example$X,
pos = gtex_example$pos, L = 15, L_greedy = 5,
verbose = TRUE)
#> HINT: ncol(Y) is not 2^J or positions are unevenly spaced; interpolated to a regular dyadic grid.
#> iter ELBO delta sigma2 mem V extras
#> 1 -205997.7982 - [0.998, 0.998, 0.999] 0.21 GB [1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] pi_null=[1.00, 1.00]
#> iter 2: max|dPIP|=4.80e-06, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] -- converged [mem: 0.21 GB]
#> [L_greedy] 1 round, greedy_lbf_cutoff=0.100, final L=5
#> round L min(lbf) action
#> 1 5 0.000 saturated
fit_s <- mf_post_smooth(fit, method = "TI")
mfsusie_plot(fit_s)
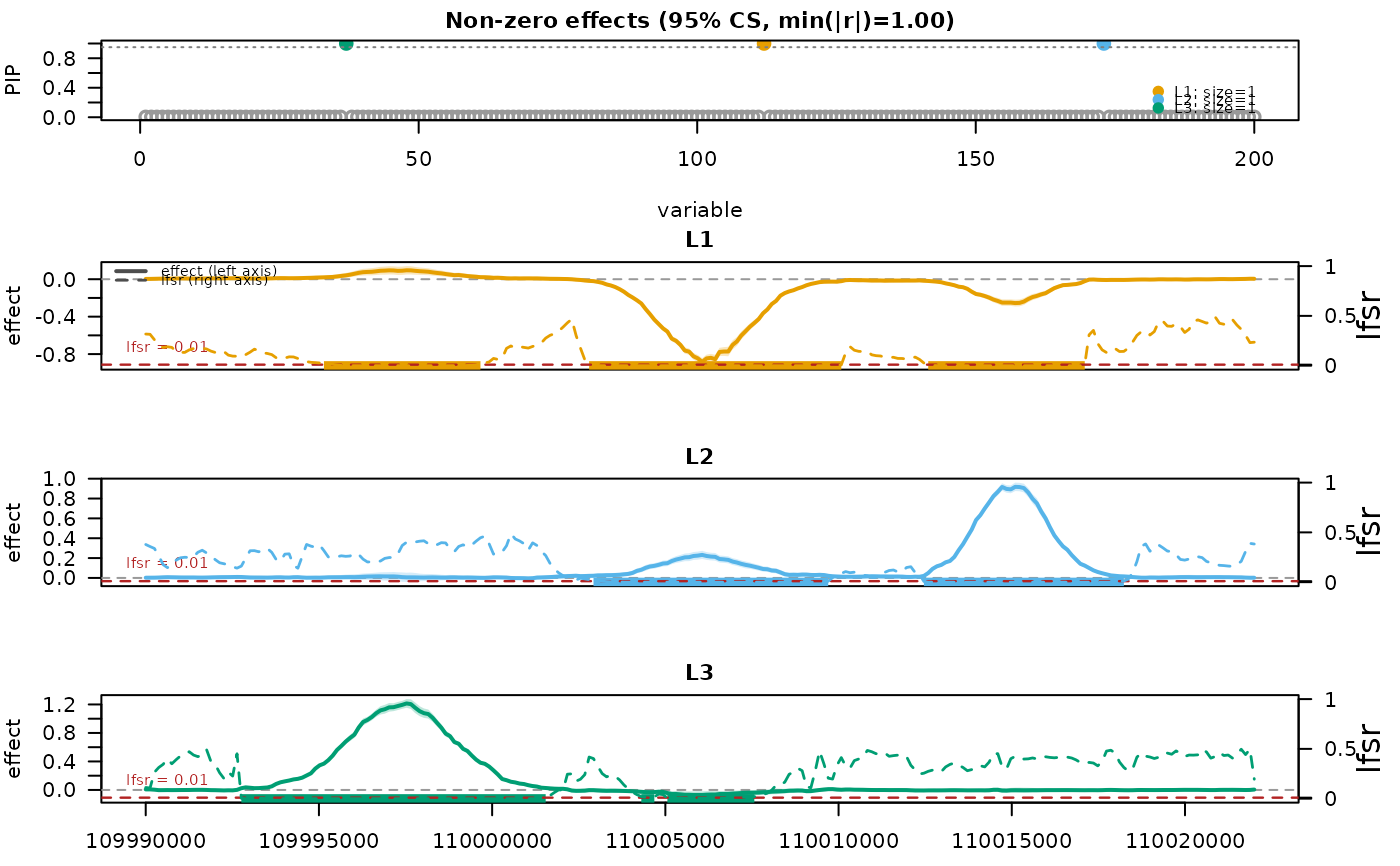 # }
# }