Simulated multi-outcome QTL data for mfsusie() covering
DNA methylation (functional, T = 64), RNA-seq (functional,
T = 32), and two scalar QTLs (eQTL, pQTL). Two causal SNPs
shared across all four outcomes; per-outcome shapes and
signs differ.
Format
A list with components
Xn x pgenotype matrix (p = 150) sliced fromsusieR::N3finemapping$X.Y_listnamed list of four outcomes:
dnam(n x 64),rna(n x 32),eqtl(n x 1),pqtl(n x 1).pos_listnamed list of position vectors (CpG bp, exon-body indices, scalar dummy
1Lfor the QTLs).causal_snpsinteger vector of shared causal SNPs.
descriptionfree-text description.
Examples
# \donttest{
data(multiomic_example)
fit <- mfsusie(multiomic_example$X, multiomic_example$Y_list,
pos = multiomic_example$pos_list, L = 15, L_greedy = 5,
verbose = TRUE)
#> HINT: ncol(Y) is not 2^J or positions are unevenly spaced; interpolated to a regular dyadic grid.
#> iter ELBO delta sigma2 mem V extras
#> 1 -78950.9731 - [0.998, 0.999, 1.000] 0.19 GB [1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] pi_null=[1.00, 1.00]
#> iter 2: max|dPIP|=1.95e-02, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.19 GB]
#> iter 3: max|dPIP|=6.97e-03, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.19 GB]
#> iter 4: max|dPIP|=6.53e-03, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.19 GB]
#> iter 5: max|dPIP|=7.53e-03, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.19 GB]
#> iter 6: max|dPIP|=1.10e-02, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.19 GB]
#> iter 7: max|dPIP|=1.72e-02, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.19 GB]
#> iter 8: max|dPIP|=1.89e-02, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.19 GB]
#> iter 9: max|dPIP|=2.90e-02, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] -- converged (stalled) [mem: 0.19 GB]
#> WARNING: PIP convergence stalled (no improvement in 5 iterations); returning current state.
#> [L_greedy] 1 round, greedy_lbf_cutoff=0.100, final L=5
#> round L min(lbf) action
#> 1 5 0.000 saturated
fit_s <- mf_post_smooth(fit, method = "TI")
#> HINT: method = 'TI' is a wavelet smoother and adds no power for outcome 3 (T_m = 1, scalar). Falling back to method = 'scalewise' for that outcome.
#> HINT: method = 'TI' is a wavelet smoother and adds no power for outcome 4 (T_m = 1, scalar). Falling back to method = 'scalewise' for that outcome.
mfsusie_plot(fit_s)
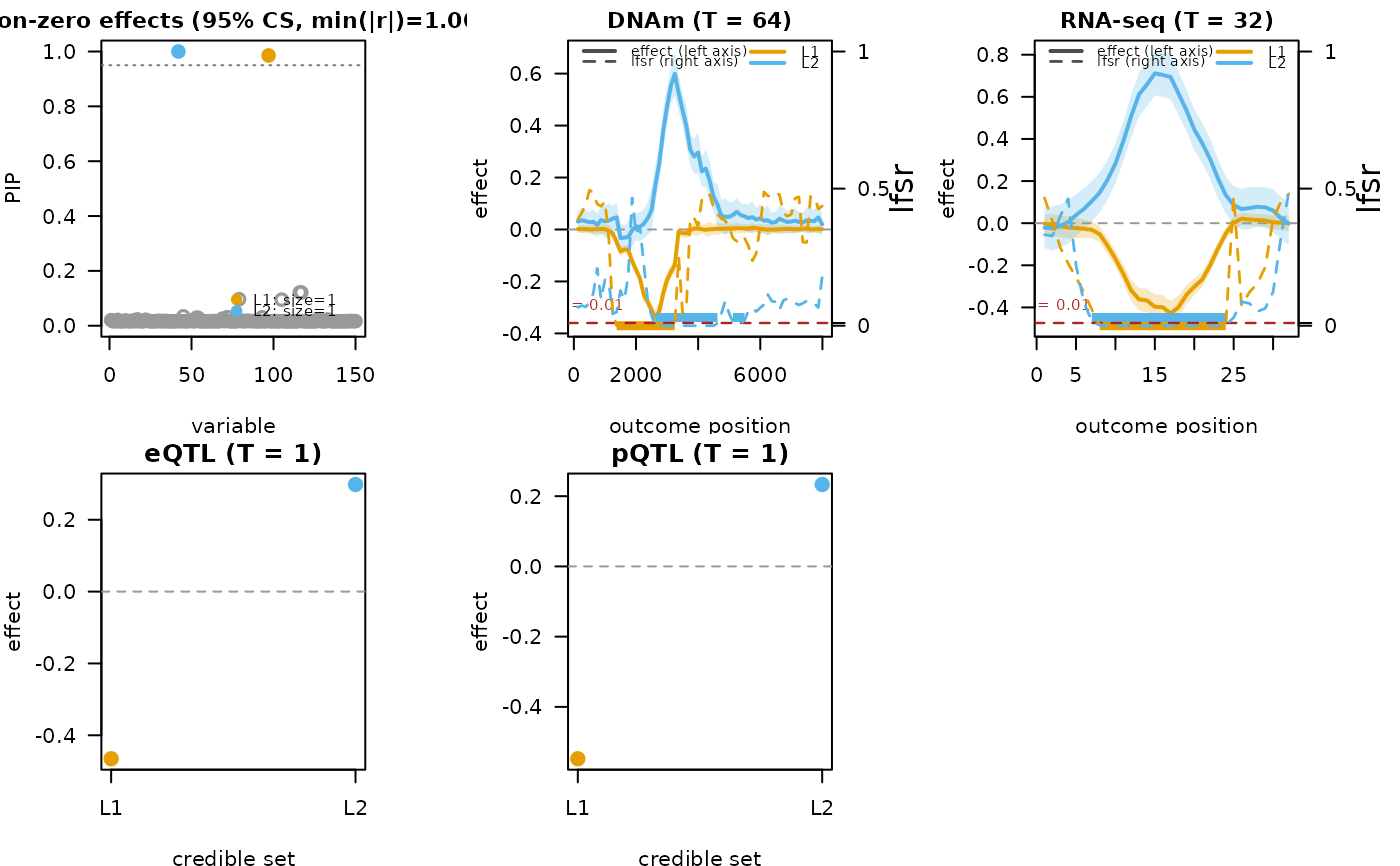 # }
# }