Simulated cis-mQTL data with n = 100 samples, p = 12
SNPs, and T = 32 CpGs. Three causal SNPs (1, 9, 3) act on
two CpG clusters: SNPs 1 and 9 affect cluster A (CpGs 9 to
16) with opposite signs, and SNP 3 affects cluster B (CpGs
25 to 32). SNP 4 is a high-LD near-clone of SNP 3.
Format
A list with components
Xn x pgenotype matrix (p = 12).Yn x Tmethylation matrix (T = 32).poslength-
Tinteger vector of CpG indices.causal_snpsinteger vector
c(1, 9, 3)of column indices inXthat carry true effects.causal_betaslength(causal_snps) x Tmatrix of true per-CpG effects.truth_masklength-3 list of length-
Tboolean vectors, one per CS, marking the CpGs each CS truly affects. Read bymfsusie_plot_lfsr()when thetruthargument is supplied.descriptionfree-text description of the dataset.
Examples
# \donttest{
data(dnam_example)
fit <- fsusie(dnam_example$Y, dnam_example$X,
pos = dnam_example$pos, L = 15, L_greedy = 5, verbose = TRUE)
#> iter ELBO delta sigma2 mem V extras
#> 1 -4503.3135 - [0.988, 0.991, 0.997] 0.18 GB [1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] pi_null=[1.00, 1.00]
#> iter 2: max|dPIP|=5.91e-01, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.18 GB]
#> iter 3: max|dPIP|=3.26e-01, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.18 GB]
#> iter 4: max|dPIP|=5.75e-03, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.18 GB]
#> iter 5: max|dPIP|=9.71e-03, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] [mem: 0.18 GB]
#> iter 6: max|dPIP|=1.35e-05, V=[1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00, 1.00e+00] -- converged [mem: 0.18 GB]
#> [L_greedy] 1 round, greedy_lbf_cutoff=0.100, final L=5
#> round L min(lbf) action
#> 1 5 0.000 saturated
fit_s <- mf_post_smooth(fit, method = "TI")
mfsusie_plot(fit_s)
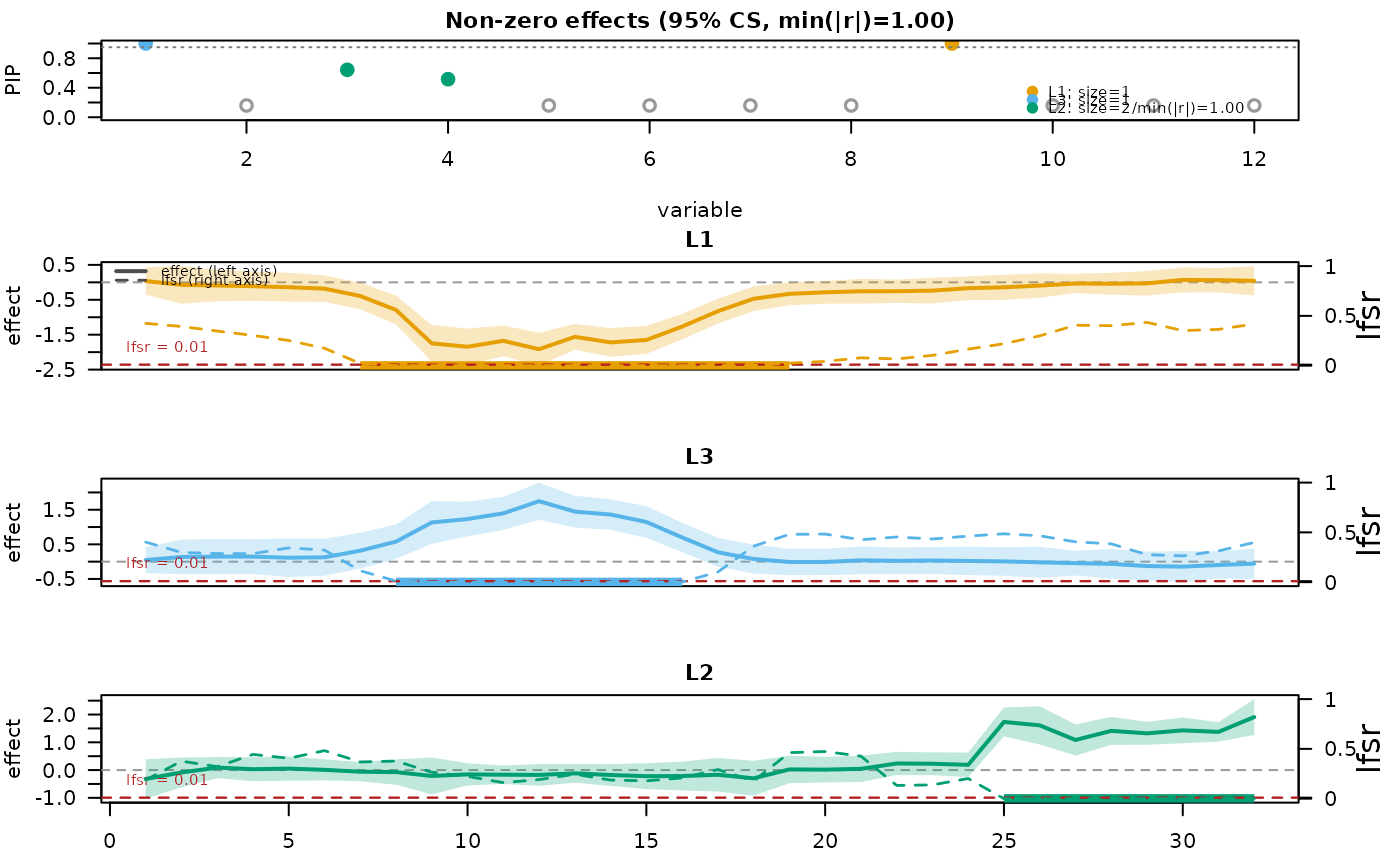 # }
# }